RocR
- Description: transcriptional activator of arginine utilization operons
| Gene name | rocR |
| Synonyms | |
| Essential | no |
| Product | transcriptional regulator |
| Function | transcriptional activator of arginine utilization operons |
| MW, pI | 52.6 kDa, 6.08 |
| Gene length, protein length | 1383 bp, 461 amino acids |
| Immediate neighbours | rocD, yyxA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 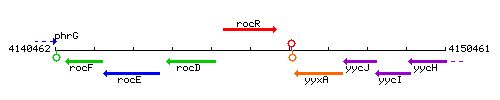
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU40350
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transcription activation at SigL-dependent promoters of rocABC, rocDEF, and rocG
- Protein family: Sigma-54 interacting transcription activator
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Sigma-54 factor interaction domain (143–372)
- 2x nucleotid-binding domain (171–178),(233–242)
- HTH motif (434–453)
- Modification:
- Cofactor(s): ornithine or citrulline are required for RocR-dependent transcription activation PubMed
- Effectors of protein activity:
- Interactions: RocR-SigL
- Localization:
Database entries
- Structure:
- Swiss prot entry: P38022
- KEGG entry: [3]
Additional information
Expression and regulation
- Operon: rocR
- Regulatory mechanism: autorepression
- Additional information:
Biological materials
- Mutant: QB5533 (aphA3), from Michel Debarbouille, available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- Antibody:
Labs working on this gene/protein
Michel Debarbouille, Pasteur Institute, Paris, France Homepage
Your additional remarks
References
- Belitsky BR, Sonenshein AL (1998) Role and regulation of Bacillus subtilis glutamate dehydrogenase genes. J Bacteriol 180:6298-6305. PubMed
- Belitsky BR, Sonenshein, AL: (1999) An enhancer element located downstream of the major glutamate dehydrogenase gene of Bacillus subtilis. Proc Natl Acad Sci USA , 96:10290-10295. PubMed
- Gardan R, Rapoport G, Débarbouillé M:(1997) Role of the transcriptional activator RocR in the arginine-degradation pathway of Bacillus subtilis. Mol Microbiol , 24:825-837. PubMed
- Gardan R, Rapoport G, Débarbouillé M: (1995) Expression of the rocDEF operon involved in arginine catabolism in Bacillus subtilis. J Mol Biol , 249:843-856. PubMed
- Ali, N. O., J. Jeusset, E. Larquet, E. le Cam, B. Belitsky, A. L. Sonenshein, T. Msadek, and M. Débarbouillé. 2003. Specificity of the interaction of RocR with the rocG-rocA intergenic region in Bacillus subtilis. Microbiology 149: 739-750. PubMed
- Calogero S, Gardan R, Glaser P, Schweizer J, Rapoport G, Debarbouille M. (1994) RocR, a novel regulatory protein controlling arginine utilization in Bacillus subtilis, belongs to the NtrC/NifA family of transcriptional activators. J. Bacteriol. volume: page-page. PubMed
- Author1, Author2 & Author3 (year) Title # Author1, Author2 & Author3 (year) Title Journal 176: 1234-1241. PubMed