Rny
- Description: RNase Y, 5' end sensitive endoribonuclease, involved in the degradation/processing of mRNA
| Gene name | rny |
| Synonyms | ymdA |
| Essential | yes |
| Product | RNase Y |
| Function | RNA processing and degradation |
| Gene expression levels in SubtiExpress: rny | |
| Interactions involving this protein in SubtInteract: Rny | |
| Regulatory function of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 58,7 kDa, 5.39 |
| Gene length, protein length | 1560 bp, 520 amino acids |
| Immediate neighbours | pbpX, ymdB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 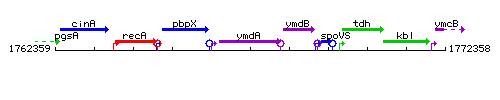
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed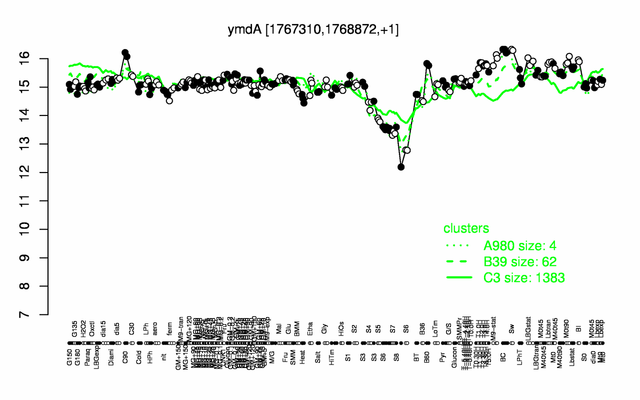
| |
Contents
Categories containing this gene/protein
Rnases, biofilm formation, essential genes, membrane proteins
This gene is a member of the following regulons
Targets of RNase Y
The gene
Basic information
- Locus tag: BSU16960
Phenotypes of a mutant
- essential PubMed
- transcription profile resulting from rny depletion: GEO PubMed
- defect in spore germination PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- RNase Y cleaves S-box riboswitch RNAs in vivo and in vitro PubMed
- preference for 5' monophosphorylated substrate in vitro PubMed
- endonucleolytic cleavage PubMed
- required for the processing of the gapA operon mRNA PubMed
- cleavage activity appears sensitive to downstream secondary structure PubMed
- RNase Y initiates the degradation of rpsO mRNA PubMed
- RNase Y is responsible for the degradation of 23S rRNA, 16S rRNA, and mRNAs in aging spores PubMed
- Protein family: Member of the HD superfamily of metal-dependent phosphohydrolases; 2',3' cyclic nucleotide phosphodiesterase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): requires Mg+2, which can be replaced by Zn+2 or Mn+2 ions, PubMed
- Effectors of protein activity: appears sensitive to downstream secondary structure, PubMed
Database entries
- Structure:
- UniProt: O31774
- KEGG entry: [3]
- E.C. number: 3.1.4.16
Additional information
required for the processing of the gapA operon mRNA
Expression and regulation
- Sigma factor:
- Regulation: constitutive
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- essential!!!!
- 4043 (rny under p-spac control, cat), GP193 (rny under p-xyl control, cat), both available in Jörg Stülke's lab
- SSB447 (rny under P-spac control, "erm") available in Putzer lab.
- Expression vector:
- N-terminal Strep-tag, expression in E. coli, in pGP172: pGP441, available in Jörg Stülke's lab
- N-terminal Strep-tag, for SPINE, expression in B. subtilis, in pGP380: pGP775, available in Jörg Stülke's lab
- C-terminal Strep-tag, for SPINE, expression in B. subtilis, in pGP382: pGP1852, available in Jörg Stülke's lab
- Expression of RNase Y missing the N-terminal transmembrane domain (25aa) as an intein fusion in E. coli (no tag left in the purified protein) available in the Putzer lab
- wild type rny, expression in B. subtilis, in pBQ200: pGP1201, available in Jörg Stülke's lab
- there is also a series of domain constructs present in pBQ200, all available in Jörg Stülke's lab
- chromosomal expression of Rny-Strep, spc: GP1033, available in Jörg Stülke's lab
- lacZ fusion: pGP459 (in pAC7), available in Jörg Stülke's lab
- GFP fusion:
- B. subtilis 3569 (amyE:: (p-xyl rny-gfpmut1-spc)), available in Errington lab
- pGP1368 for chromosomal expression of rny-YFP, available in Jörg Stülke's lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- FLAG-tag construct: GP1030 (spc, based on pGP1331), available in Jörg Stülke's lab
- Antibody: available in van Dijl and in Jörg Stülke's lab
Labs working on this gene/protein
Harald Putzer, IBPC Paris, France Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Additional reviews: PubMed
Lehnik-Habrink M, Lewis RJ, Mäder U, Stülke J RNA degradation in Bacillus subtilis: an interplay of essential endo- and exoribonucleases. Mol Microbiol.: 2012, 84(6) 1005-1017. PubMed:22568516
Publications on B. subtilis rny
Additional publications: PubMed
Diethmaier C, Pietack N, Gunka K, Wrede C, Lehnik-Habrink M, Herzberg C, Hübner S, Stülke J A Novel Factor Controlling Bistability in Bacillus subtilis: The YmdB Protein Affects Flagellin Expression and Biofilm Formation. J Bacteriol.: 2011, 193(21):5997-6007. PubMed:21856853
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Publications on homologs from other organisms
Song Ok Kang, Michael G Caparon, Kyu Hong Cho
Virulence gene regulation by CvfA, a putative RNase: the CvfA-enolase complex in Streptococcus pyogenes links nutritional stress, growth-phase control, and virulence gene expression.
Infect Immun: 2010, 78(6);2754-67
[PubMed:20385762]
[WorldCat.org]
[DOI]
(I p)
Makiko Nagata, Chikara Kaito, Kazuhisa Sekimizu
Phosphodiesterase activity of CvfA is required for virulence in Staphylococcus aureus.
J Biol Chem: 2008, 283(4);2176-84
[PubMed:17951247]
[WorldCat.org]
[DOI]
(P p)
Chikara Kaito, Kenji Kurokawa, Yasuhiko Matsumoto, Yutaka Terao, Shigetada Kawabata, Shigeyuki Hamada, Kazuhisa Sekimizu
Silkworm pathogenic bacteria infection model for identification of novel virulence genes.
Mol Microbiol: 2005, 56(4);934-44
[PubMed:15853881]
[WorldCat.org]
[DOI]
(P p)