Eno
- Description: enolase, glycolytic/ gluconeogenic enzyme, universally conserved protein
| Gene name | eno |
| Synonyms | |
| Essential | no |
| Product | enolase |
| Function | enzyme in glycolysis/ gluconeogenesis |
| Interactions involving this protein in SubtInteract: Eno | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 46,4 kDa, 4.49 |
| Gene length, protein length | 1290 bp, 430 amino acids |
| Immediate neighbours | yvbK, pgm |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 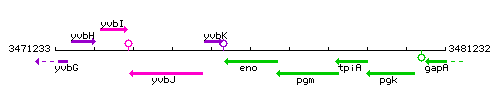
This image was kindly provided by SubtiList
| |
Expression 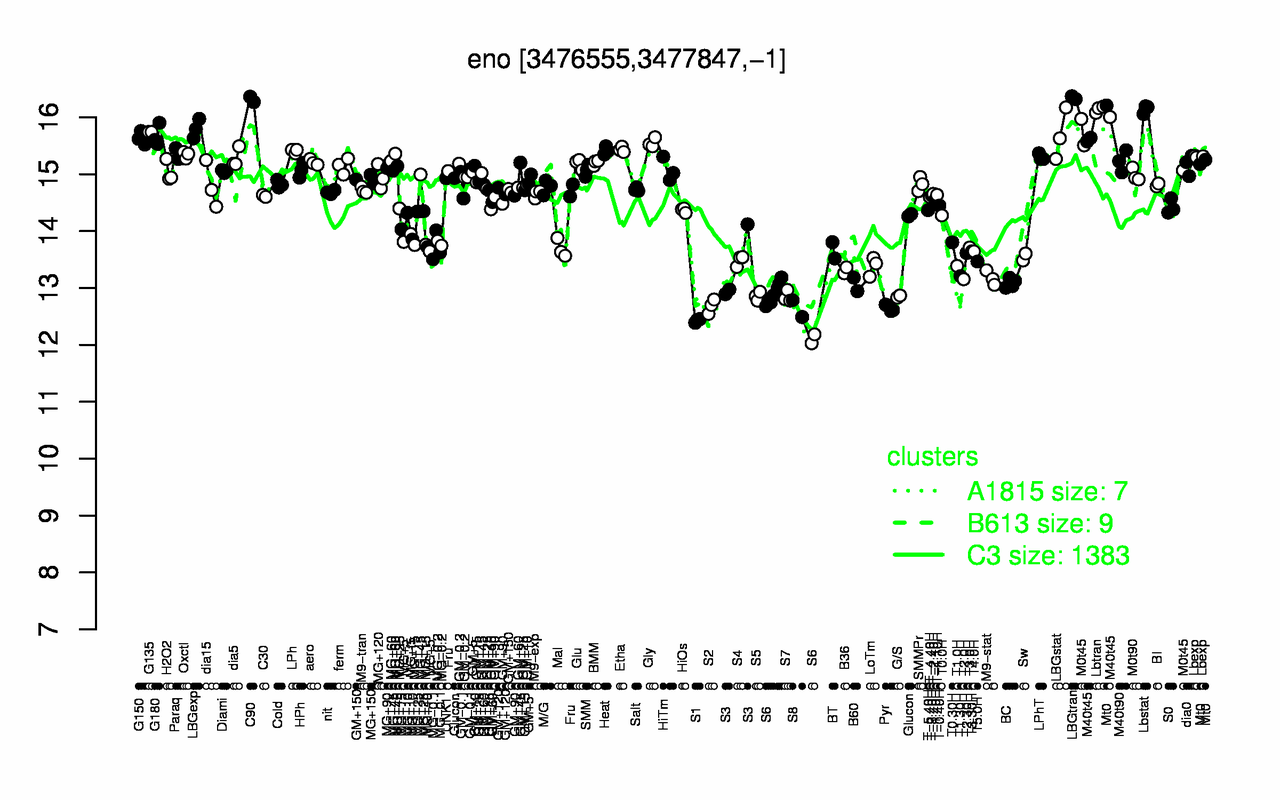
| |
Contents
Categories containing this gene/protein
carbon core metabolism, membrane proteins, phosphoproteins, universally conserved proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU33900
Phenotypes of a mutant
- no growth on LB, requires glucose and malate
- essential according to Kobayashi et al. on LB PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2-phospho-D-glycerate = phosphoenolpyruvate + H2O (according to Swiss-Prot) 2-phospho-D-glycerate = phosphoenolpyruvate + H(2)O
- Protein family: enolase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: reversible Michaelis-Menten PubMed
- Domains:
- substrate binding domain (366–369)
- Cofactor(s): Mg2+
- Effectors of protein activity:
- Inhibited by EDTA PubMed
Database entries
- UniProt: P37869
- KEGG entry: [3]
- E.C. number: 4.2.1.11
Additional information
- Enolase is a moonlighting protein. PubMed
- There are indications that this enzyme is an octamer PubMed
- universally conserved protein
- extensive information on the structure and enzymatic properties of Eno can be found at Proteopedia
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- pGP1426 (expression of eno in B. subtilis, in pBQ200), available in Stülke lab
- pGP1500 (expression of pgm and eno in B. subtilis, in pBQ200), available in Stülke lab
- pGP563 (N-terminal His-tag, in pWH844), available in Stülke lab
- pGP1276 (N-terminal Strep-tag, purification from E. coli, in pGP172), available in Stülke lab
- pGP93 (N-terminal Strep-tag, purification from B. subtilis, for SPINE, in pGP380), available in Stülke lab
- GP1215 (eno-Strep (spc)), purification from B. subtilis, for SPINE, available in Stülke lab
- lacZ fusion:
- see pgk
- GFP fusion: pHT315-yfp-eno, available in Mijakovic lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
G H Reed, R R Poyner, T M Larsen, J E Wedekind, I Rayment
Structural and mechanistic studies of enolase.
Curr Opin Struct Biol: 1996, 6(6);736-43
[PubMed:8994873]
[WorldCat.org]
[DOI]
(P p)
Subcellular localization of enolase
Additional publications: PubMed
Carsten Jers, Malene Mejer Pedersen, Dafni Katerina Paspaliari, Wolfgang Schütz, Christina Johnsson, Boumediene Soufi, Boris Macek, Peter Ruhdal Jensen, Ivan Mijakovic
Bacillus subtilis BY-kinase PtkA controls enzyme activity and localization of its protein substrates.
Mol Microbiol: 2010, 77(2);287-99
[PubMed:20497499]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Grégory Boël, Vianney Pichereau, Ivan Mijakovic, Alain Mazé, Sandrine Poncet, Sylvie Gillet, Jean-Christophe Giard, Axel Hartke, Yanick Auffray, Josef Deutscher
Is 2-phosphoglycerate-dependent automodification of bacterial enolases implicated in their export?
J Mol Biol: 2004, 337(2);485-96
[PubMed:15003462]
[WorldCat.org]
[DOI]
(P p)
Other original publications