Difference between revisions of "SrfAC"
(→Original publications) |
|||
| Line 154: | Line 154: | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>17227471,16166527,8288534,18583577, 17190806,8830686,17218307, 1715856 12642660 16091051 18763711 22511326,20817675</pubmed> | + | <pubmed>17227471,16166527,8288534,18583577, 17190806,8830686,17218307, 1715856 12642660 16091051 18763711 23957147 22511326,20817675</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 07:33, 21 August 2013
- Description: surfactin synthetase / competence
| Gene name | srfAC |
| Synonyms | comL |
| Essential | no |
| Product | surfactin synthetase / competence |
| Function | antibiotic synthesis |
| Gene expression levels in SubtiExpress: srfAC | |
| MW, pI | 143 kDa, 4.97 |
| Gene length, protein length | 3822 bp, 1274 aa |
| Immediate neighbours | comS, srfAD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 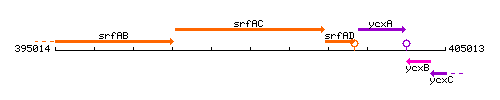
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed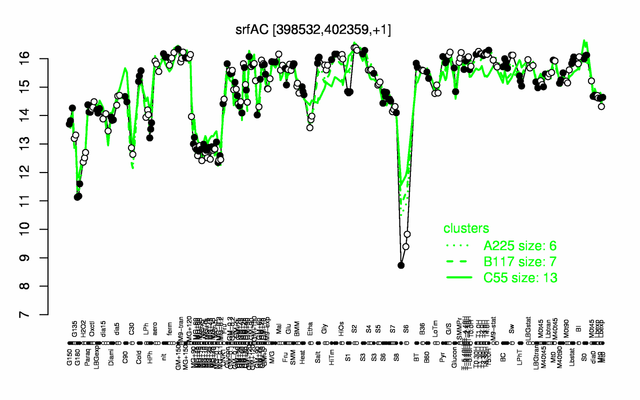
| |
Contents
Categories containing this gene/protein
miscellaneous metabolic pathways, biosynthesis of antibacterial compounds, membrane proteins, phosphoproteins
This gene is a member of the following regulons
Abh regulon, CodY regulon, ComA regulon, PerR regulon, Spx regulon
The gene
Basic information
- Locus tag: BSU03510
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: acyl carrier domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on Ser-1003 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane PubMed
Database entries
- Structure: 2VSQ
- UniProt: Q08787
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications