Difference between revisions of "TapA"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
(→Extended information on the protein) |
||
| Line 88: | Line 88: | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
** [[TapA]]-[[TasA]] {{PubMed|21477127}} | ** [[TapA]]-[[TasA]] {{PubMed|21477127}} | ||
| + | ** [[SipW]]-[[TapA]] {{PubMed|10559173}} | ||
* '''[[Localization]]:''' | * '''[[Localization]]:''' | ||
** attached to the cell surface (on the outside of the cell), associated with peptidoglycan {{PubMed|21477127}} | ** attached to the cell surface (on the outside of the cell), associated with peptidoglycan {{PubMed|21477127}} | ||
| − | ** secretion requires SipW | + | ** secretion requires [[SipW]] {{PubMed|10559173}} |
=== Database entries === | === Database entries === | ||
Revision as of 09:28, 23 March 2012
- Description: required for the anchoring of the TasA amyloid fibers to the cell and for the initiation of fiber polymerization, minor fiber component
| Gene name | tapA |
| Synonyms | yqhD, yqxM |
| Essential | no |
| Product | TasA anchoring/assembly protein |
| Function | biofilm formation |
| Interactions involving this protein in SubtInteract: TapA | |
| Regulation of this protein in SubtiPathways: Biofilm, Protein secretion | |
| MW, pI | 28 kDa, 6.677 |
| Gene length, protein length | 759 bp, 253 aa |
| Immediate neighbours | sipW, yqzG |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 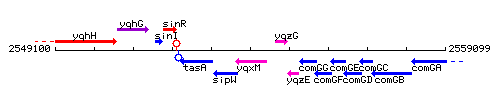
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
biofilm formation, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU24640
Phenotypes of a mutant
The mutants are able to form a biofilm in the presence of D-amino acids PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity: D-amino acids lead to disappearance of TapA from the cell wall PubMed
Database entries
- Structure:
- UniProt: P40949
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Diethmaier C, Pietack N, Gunka K, Wrede C, Lehnik-Habrink M, Herzberg C, Hübner S, Stülke J A Novel Factor Controlling Bistability in Bacillus subtilis: The YmdB Protein Affects Flagellin Expression and Biofilm Formation. J Bacteriol.: 2011, 193(21):5997-6007. PubMed:21856853
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Martin Lehnik-Habrink, Joseph Newman, Fabian M Rothe, Alexandra S Solovyova, Cecilia Rodrigues, Christina Herzberg, Fabian M Commichau, Richard J Lewis, Jörg Stülke
RNase Y in Bacillus subtilis: a Natively disordered protein that is the functional equivalent of RNase E from Escherichia coli.
J Bacteriol: 2011, 193(19);5431-41
[PubMed:21803996]
[WorldCat.org]
[DOI]
(I p)
Martin Lehnik-Habrink, Henrike Pförtner, Leonie Rempeters, Nico Pietack, Christina Herzberg, Jörg Stülke
The RNA degradosome in Bacillus subtilis: identification of CshA as the major RNA helicase in the multiprotein complex.
Mol Microbiol: 2010, 77(4);958-71
[PubMed:20572937]
[WorldCat.org]
[DOI]
(I p)
Irnov Irnov, Cynthia M Sharma, Jörg Vogel, Wade C Winkler
Identification of regulatory RNAs in Bacillus subtilis.
Nucleic Acids Res: 2010, 38(19);6637-51
[PubMed:20525796]
[WorldCat.org]
[DOI]
(I p)
Shiyi Yao, David H Bechhofer
Initiation of decay of Bacillus subtilis rpsO mRNA by endoribonuclease RNase Y.
J Bacteriol: 2010, 192(13);3279-86
[PubMed:20418391]
[WorldCat.org]
[DOI]
(I p)
Jessica C Zweers, Thomas Wiegert, Jan Maarten van Dijl
Stress-responsive systems set specific limits to the overproduction of membrane proteins in Bacillus subtilis.
Appl Environ Microbiol: 2009, 75(23);7356-64
[PubMed:19820159]
[WorldCat.org]
[DOI]
(I p)
Karen Shahbabian, Ailar Jamalli, Léna Zig, Harald Putzer
RNase Y, a novel endoribonuclease, initiates riboswitch turnover in Bacillus subtilis.
EMBO J: 2009, 28(22);3523-33
[PubMed:19779461]
[WorldCat.org]
[DOI]
(I p)
Fabian M Commichau, Fabian M Rothe, Christina Herzberg, Eva Wagner, Daniel Hellwig, Martin Lehnik-Habrink, Elke Hammer, Uwe Völker, Jörg Stülke
Novel activities of glycolytic enzymes in Bacillus subtilis: interactions with essential proteins involved in mRNA processing.
Mol Cell Proteomics: 2009, 8(6);1350-60
[PubMed:19193632]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Alison Hunt, Joy P Rawlins, Helena B Thomaides, Jeff Errington
Functional analysis of 11 putative essential genes in Bacillus subtilis.
Microbiology (Reading): 2006, 152(Pt 10);2895-2907
[PubMed:17005971]
[WorldCat.org]
[DOI]
(P p)