Difference between revisions of "ClpE"
(→References) |
(→References) |
||
| Line 136: | Line 136: | ||
=References= | =References= | ||
| + | ==Reviews== | ||
| + | <pubmed> 17302811 </pubmed> | ||
| + | ==Original Publications== | ||
'''Additional publications:''' {{PubMed|16163393,17380125}} | '''Additional publications:''' {{PubMed|16163393,17380125}} | ||
<pubmed>10320580 9987115 11179229 21821766</pubmed> | <pubmed>10320580 9987115 11179229 21821766</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 09:40, 31 October 2011
- Description: ATP-dependent Clp protease-like (class III stress gene)
| Gene name | clpE |
| Synonyms | ykvH |
| Essential | no |
| Product | ATP-dependent Clp protease-like |
| Function | protein degradation |
| Interactions involving this protein in SubtInteract: ClpE | |
| Metabolic function and regulation of this protein in SubtiPathways: Stress | |
| MW, pI | 77 kDa, 5.135 |
| Gene length, protein length | 2097 bp, 699 aa |
| Immediate neighbours | motA, ykvI |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 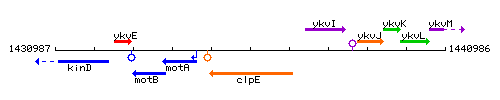
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
proteolysis, heat shock proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13700
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATPase/chaperone
Extended information on the protein
- Kinetic information:
- Domains: AAA-ATPase PFAM
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O31673
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: clpE PubMed
- Additional information:
Biological materials
- Mutant: clpE::spec available from the Hamoen] Lab
- Expression vector:
- lacZ fusion:
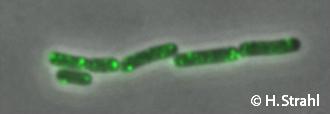
- GFP fusion: C-terminal YFP and CFP fusions (single copy) available from the Hamoen] Lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Newcastle University, UK homepage
Your additional remarks
References
Reviews
Original Publications
Additional publications: PubMed