Difference between revisions of "RnjA"
(→RNAs affected by rnjA) |
(→Original publications) |
||
| Line 161: | Line 161: | ||
==Original publications== | ==Original publications== | ||
'''Additional publications:''' {{PubMed|21893285,21893286,21908660}} | '''Additional publications:''' {{PubMed|21893285,21893286,21908660}} | ||
| − | <pubmed>18079181, 19553197, 17981983, 19458035, 15831787, 18204464, 18713320, 19193632, 17005971, 18445592, 17512403, 17229210, 17576666, 19210617 19633085 19638340 19850915 19880604 20025672 20418391 20572937 ,21803996 21862575 </pubmed> | + | <pubmed>18079181, 19553197, 17981983, 19458035, 15831787, 18204464, 18713320, 19193632, 17005971, 18445592, 17512403, 17229210, 17576666, 19210617 19633085 19638340 19850915 19880604 20025672 20418391 20572937 ,21803996 21862575 21925382 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 16:05, 20 September 2011
- Description: RNase J1
| Gene name | rnjA |
| Synonyms | ykqC |
| Essential | yes PubMed |
| Product | RNase J1 |
| Function | RNA processing |
| Interactions involving this protein in SubtInteract: RNase J1 | |
| Metabolic function and regulation of this protein in SubtiPathways: Ile, Leu, Val, Coenzyme A | |
| MW, pI | 61 kDa, 5.902 |
| Gene length, protein length | 1665 bp, 555 aa |
| Immediate neighbours | adeC, ykzG |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 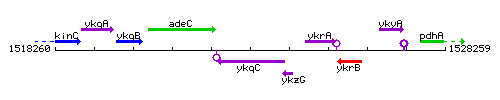
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU14530
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: endonuclease and 5'-3' exonuclease
- Protein family: RNase J subfamily (according to Swiss-Prot)
- Paralogous protein(s): RnjB
RNAs affected by rnjA
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: 3BK1 (RNase J from Thermus thermophilus) 3BK2 (RNase J from Thermus thermophilus, complex with UMP)
- UniProt: Q45493
- KEGG entry: [2]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
- required for thrS RNA processing, involved in maturation of the 5’-end of the16S rRNA
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant: GP41 (rnjA under control of p(xyl)), available in Stülke lab; SSB342 (rnjA under pspac), cat, available in Harald Putzer lab
- Expression vector:
- for chromosomal expression of RNase J1-Strep (spc): GP1034, available in Jörg Stülke's lab
- for chromosomal expression of RNase J1-Strep (cat): GP1042, available in Jörg Stülke's lab
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Harald Putzer, IBPC Paris, France Homepage
David Bechhofer, Mount Sinai School, New York, USA Homepage
Ciaran Condon, IBPC, Paris, France Homepage
Your additional remarks
References
Reviews
Original publications
Additional publications: PubMed