Difference between revisions of "ScoC"
m (→Biological materials) |
|||
| Line 133: | Line 133: | ||
* '''Mutant:''' Strain OM213 of Ogura MA, et al {{PubMed|15126467}}, genotype ''trpC2 scoC::pMutin-scoC leuC7'', available as [http://bgsc.org BGSC] 1A918 | * '''Mutant:''' Strain OM213 of Ogura MA, et al {{PubMed|15126467}}, genotype ''trpC2 scoC::pMutin-scoC leuC7'', available as [http://bgsc.org BGSC] 1A918 | ||
| + | ** 1A918 ( ''scoC''::''erm''), {{PubMed|15126467}}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1A918&Search=1A918 BGSC] | ||
* '''Expression vector:''' for expression, purification in E. coli with N-terminal His-tag, pRSETA available in Gerth lab | * '''Expression vector:''' for expression, purification in E. coli with N-terminal His-tag, pRSETA available in Gerth lab | ||
Revision as of 10:21, 19 September 2012
- Description: transcriptional repressor of genes expressed in the transition phase
| Gene name | scoC |
| Synonyms | hpr, catA |
| Essential | no |
| Product | transcriptional repressor (MarR family) |
| Function | transition state regulator |
| Gene expression levels in SubtiExpress: scoC | |
| Interactions involving this protein in SubtInteract: ScoC | |
| Regulation of this protein in SubtiPathways: Biofilm, Alternative nitrogen sources | |
| MW, pI | 23 kDa, 5.188 |
| Gene length, protein length | 609 bp, 203 aa |
| Immediate neighbours | yhaI, yhaH |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 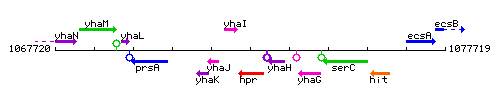
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed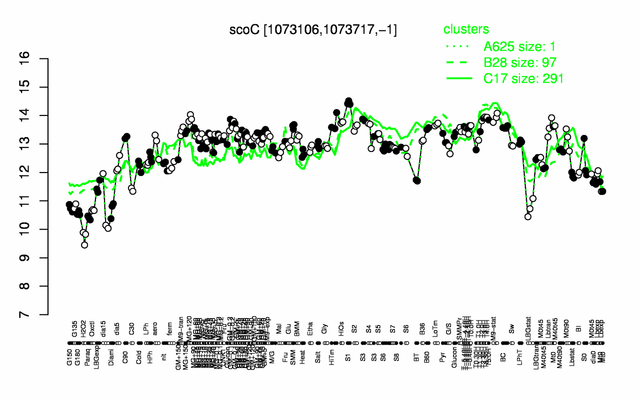
| |
Contents
Categories containing this gene/protein
transcription factors and their control, transition state regulators, phosphoproteins
This gene is a member of the following regulons
AbrB regulon, SalA regulon, SenS regulon
The ScoC regulon
The gene
Basic information
- Locus tag: BSU09990
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-3 PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 2FXA
- UniProt: P11065
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: scoC PubMed
- Sigma factor:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: Strain OM213 of Ogura MA, et al PubMed, genotype trpC2 scoC::pMutin-scoC leuC7, available as BGSC 1A918
- Expression vector: for expression, purification in E. coli with N-terminal His-tag, pRSETA available in Gerth lab
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Bindiya Kaushal, Salbi Paul, F Marion Hulett
Direct regulation of Bacillus subtilis phoPR transcription by transition state regulator ScoC.
J Bacteriol: 2010, 192(12);3103-13
[PubMed:20382764]
[WorldCat.org]
[DOI]
(I p)
Prashant Kodgire, K Krishnamurthy Rao
A dual mode of regulation of flgM by ScoC in Bacillus subtilis.
Can J Microbiol: 2009, 55(8);983-9
[PubMed:19898538]
[WorldCat.org]
[DOI]
(I p)
Takashi Inaoka, Guojun Wang, Kozo Ochi
ScoC regulates bacilysin production at the transcription level in Bacillus subtilis.
J Bacteriol: 2009, 191(23);7367-71
[PubMed:19801406]
[WorldCat.org]
[DOI]
(I p)
Prashant Kodgire, K Krishnamurthy Rao
hag expression in Bacillus subtilis is both negatively and positively regulated by ScoC.
Microbiology (Reading): 2009, 155(Pt 1);142-149
[PubMed:19118355]
[WorldCat.org]
[DOI]
(P p)
Prashant Kodgire, Madhulika Dixit, K Krishnamurthy Rao
ScoC and SinR negatively regulate epr by corepression in Bacillus subtilis.
J Bacteriol: 2006, 188(17);6425-8
[PubMed:16923912]
[WorldCat.org]
[DOI]
(P p)
Eiji Kawachi, Sadanobu Abe, Teruo Tanaka
Inhibition of Bacillus subtilis scoC expression by multicopy senS.
J Bacteriol: 2005, 187(24);8526-30
[PubMed:16321961]
[WorldCat.org]
[DOI]
(P p)
Mitsuo Ogura, Atsushi Matsuzawa, Hirofumi Yoshikawa, Teruo Tanaka
Bacillus subtilis SalA (YbaL) negatively regulates expression of scoC, which encodes the repressor for the alkaline exoprotease gene, aprE.
J Bacteriol: 2004, 186(10);3056-64
[PubMed:15126467]
[WorldCat.org]
[DOI]
(P p)
Alejandro Sánchez, Jorge Olmos
Bacillus subtilis transcriptional regulators interaction.
Biotechnol Lett: 2004, 26(5);403-7
[PubMed:15104138]
[WorldCat.org]
[DOI]
(P p)
Sasha H Shafikhani, Esperanza Núñez, Terrance Leighton
ScoC mediates catabolite repression of sporulation in Bacillus subtilis.
Curr Microbiol: 2003, 47(4);327-36
[PubMed:14629015]
[WorldCat.org]
[DOI]
(P p)
A Koide, M Perego, J A Hoch
ScoC regulates peptide transport and sporulation initiation in Bacillus subtilis.
J Bacteriol: 1999, 181(13);4114-7
[PubMed:10383984]
[WorldCat.org]
[DOI]
(P p)
P T Kallio, J E Fagelson, J A Hoch, M A Strauch
The transition state regulator Hpr of Bacillus subtilis is a DNA-binding protein.
J Biol Chem: 1991, 266(20);13411-7
[PubMed:1906467]
[WorldCat.org]
(P p)
B C Dowds, J A Hoch
Regulation of the oxidative stress response by the hpr gene in Bacillus subtilis.
J Gen Microbiol: 1991, 137(5);1121-5
[PubMed:1907636]
[WorldCat.org]
[DOI]
(P p)
M A Strauch, G B Spiegelman, M Perego, W C Johnson, D Burbulys, J A Hoch
The transition state transcription regulator abrB of Bacillus subtilis is a DNA binding protein.
EMBO J: 1989, 8(5);1615-21
[PubMed:2504584]
[WorldCat.org]
[DOI]
(P p)
M Perego, J A Hoch
Sequence analysis and regulation of the hpr locus, a regulatory gene for protease production and sporulation in Bacillus subtilis.
J Bacteriol: 1988, 170(6);2560-7
[PubMed:3131303]
[WorldCat.org]
[DOI]
(P p)