Difference between revisions of "TapA"
(→Original publications) |
|||
| Line 156: | Line 156: | ||
<big>Mol Microbiol. 2011 81(6): 1459-1473. </big> | <big>Mol Microbiol. 2011 81(6): 1459-1473. </big> | ||
[http://www.ncbi.nlm.nih.gov/pubmed/21815947 PubMed:21815947] | [http://www.ncbi.nlm.nih.gov/pubmed/21815947 PubMed:21815947] | ||
| − | <pubmed>18763711,19193632,17005971 19779461 19820159 20418391 20525796 20572937,21803996 | + | <pubmed>18763711,19193632,17005971 19779461 19820159 20418391 20525796 20572937,21803996 </pubmed> |
<pubmed>18430133,18047568,18647168, 20351052,16430695,10464223,17720793, 19605685 20431016 10559173 21477127 </pubmed> | <pubmed>18430133,18047568,18647168, 20351052,16430695,10464223,17720793, 19605685 20431016 10559173 21477127 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 20:17, 19 November 2011
- Description: required for the anchoring of the TasA amyloid fibers to the cell and for the initiation of fiber polymerization, minor fiber component
| Gene name | tapA |
| Synonyms | yqhD, yqxM |
| Essential | no |
| Product | TasA anchoring/assembly protein |
| Function | biofilm formation |
| Interactions involving this protein in SubtInteract: TapA | |
| Regulation of this protein in SubtiPathways: Biofilm, Protein secretion | |
| MW, pI | 28 kDa, 6.677 |
| Gene length, protein length | 759 bp, 253 aa |
| Immediate neighbours | sipW, yqzG |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 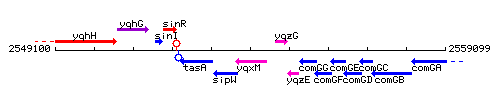
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
biofilm formation, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU24640
Phenotypes of a mutant
The mutants are able to form a biofilm in the presence of D-amino acids PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity: D-amino acids lead to disappearance of TapA from the cell wall PubMed
- Localization:
- attached to the cell surface (on the outside of the cell), associated with peptidoglycan PubMed
- secretion requires SipW
Database entries
- Structure:
- UniProt: P40949
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Diethmaier C, Pietack N, Gunka K, Wrede C, Lehnik-Habrink M, Herzberg C, Hübner S, Stülke J A Novel Factor Controlling Bistability in Bacillus subtilis: The YmdB Protein Affects Flagellin Expression and Biofilm Formation. J Bacteriol.: 2011, 193(21):5997-6007. PubMed:21856853
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Diego Romero, Hera Vlamakis, Richard Losick, Roberto Kolter
An accessory protein required for anchoring and assembly of amyloid fibres in B. subtilis biofilms.
Mol Microbiol: 2011, 80(5);1155-68
[PubMed:21477127]
[WorldCat.org]
[DOI]
(I p)
Ilana Kolodkin-Gal, Diego Romero, Shugeng Cao, Jon Clardy, Roberto Kolter, Richard Losick
D-amino acids trigger biofilm disassembly.
Science: 2010, 328(5978);627-9
[PubMed:20431016]
[WorldCat.org]
[DOI]
(I p)
Yunrong Chai, Thomas Norman, Roberto Kolter, Richard Losick
An epigenetic switch governing daughter cell separation in Bacillus subtilis.
Genes Dev: 2010, 24(8);754-65
[PubMed:20351052]
[WorldCat.org]
[DOI]
(I p)
Daniel López, Hera Vlamakis, Richard Losick, Roberto Kolter
Paracrine signaling in a bacterium.
Genes Dev: 2009, 23(14);1631-8
[PubMed:19605685]
[WorldCat.org]
[DOI]
(I p)
Kazuo Kobayashi
SlrR/SlrA controls the initiation of biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 69(6);1399-410
[PubMed:18647168]
[WorldCat.org]
[DOI]
(I p)
Frances Chu, Daniel B Kearns, Anna McLoon, Yunrong Chai, Roberto Kolter, Richard Losick
A novel regulatory protein governing biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 68(5);1117-27
[PubMed:18430133]
[WorldCat.org]
[DOI]
(I p)
Yunrong Chai, Frances Chu, Roberto Kolter, Richard Losick
Bistability and biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 67(2);254-63
[PubMed:18047568]
[WorldCat.org]
[DOI]
(P p)
Mark A Strauch, Benjamin G Bobay, John Cavanagh, Fude Yao, Angelo Wilson, Yoann Le Breton
Abh and AbrB control of Bacillus subtilis antimicrobial gene expression.
J Bacteriol: 2007, 189(21);7720-32
[PubMed:17720793]
[WorldCat.org]
[DOI]
(P p)
Frances Chu, Daniel B Kearns, Steven S Branda, Roberto Kolter, Richard Losick
Targets of the master regulator of biofilm formation in Bacillus subtilis.
Mol Microbiol: 2006, 59(4);1216-28
[PubMed:16430695]
[WorldCat.org]
[DOI]
(P p)
A G Stöver, A Driks
Control of synthesis and secretion of the Bacillus subtilis protein YqxM.
J Bacteriol: 1999, 181(22);7065-9
[PubMed:10559173]
[WorldCat.org]
[DOI]
(P p)
A G Stöver, A Driks
Regulation of synthesis of the Bacillus subtilis transition-phase, spore-associated antibacterial protein TasA.
J Bacteriol: 1999, 181(17);5476-81
[PubMed:10464223]
[WorldCat.org]
[DOI]
(P p)