Difference between revisions of "ClpP"
(→Original Publications) |
(→Reviews) |
||
| Line 131: | Line 131: | ||
==Reviews== | ==Reviews== | ||
| − | <pubmed> 19609260 </pubmed> | + | <pubmed> 19609260 19781636 </pubmed> |
| + | |||
==Original Publications== | ==Original Publications== | ||
<pubmed>17380125,9643546,10809708,11807061,9987115,14679237,18689476,15317791,17586624,11684022,12923101,17560370,12598648,16899079,19226326,9890793, 20070525,9987115,11544224 14763982 9643546, 19767395 11112444 9535081 18689473 20049702 </pubmed> | <pubmed>17380125,9643546,10809708,11807061,9987115,14679237,18689476,15317791,17586624,11684022,12923101,17560370,12598648,16899079,19226326,9890793, 20070525,9987115,11544224 14763982 9643546, 19767395 11112444 9535081 18689473 20049702 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 18:00, 3 February 2010
- Description: ATP-dependent Clp protease proteolytic subunit (class III heat-shock protein)
| Gene name | clpP |
| Synonyms | yvdN |
| Essential | no |
| Product | ATP-dependent Clp protease proteolytic subunit |
| Function | protein degradation |
| Metabolic function and regulation of this protein in SubtiPathways: Stress | |
| MW, pI | 21 kDa, 5.008 |
| Gene length, protein length | 591 bp, 197 aa |
| Immediate neighbours | trnQ-Arg, pgcM |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 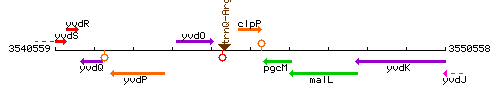
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU34540
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Hydrolysis of proteins to small peptides in the presence of ATP and magnesium (according to Swiss-Prot) endopeptidase/proteolysis
- Protein family: peptidase S14 family (according to Swiss-Prot) ClpP (IPR001907) InterPro, (PF00574) PFAM
- Paralogous protein(s):
Targets of ClpC-ClpP-dependent protein degradation
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
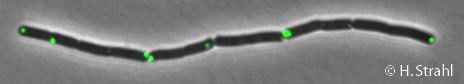
- Localization: cytoplasmic polar clusters, excluded from the nucleoid, induced clustering upon heat shock, colocalization with ClpX, ClpC and ClpE PubMed
Database entries
- Structure: Two homologue structures resolved 1TYF, 1Y7O, structural model of B. subtilis ClpP available from hstrahl
- UniProt: P80244
- KEGG entry: [3]
- E.C. number: 3.4.21.92
Additional information
Expression and regulation
- Operon: clpP PubMed
- Additional information:
Biological materials
- Mutant:
- clpP::spec and clpP::cat available , available in the Leendert Hamoen lab
- GP551 (spc), available in the Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion: C-terminal GFP fusions (both single copy and as 2th copy in amyE locus, also as CFP and YFP variants) available in the Leendert Hamoen lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Newcastle University, UK homepage
Your additional remarks
References
Reviews
Original Publications
Mitsuo Ogura, Kensuke Tsukahara
Autoregulation of the Bacillus subtilis response regulator gene degU is coupled with the proteolysis of DegU-P by ClpCP.
Mol Microbiol: 2010, 75(5);1244-59
[PubMed:20070525]
[WorldCat.org]
[DOI]
(I p)
Janine Kirstein, Anja Hoffmann, Hauke Lilie, Ronny Schmidt, Helga Rübsamen-Waigmann, Heike Brötz-Oesterhelt, Axel Mogk, Kürşad Turgay
The antibiotic ADEP reprogrammes ClpP, switching it from a regulated to an uncontrolled protease.
EMBO Mol Med: 2009, 1(1);37-49
[PubMed:20049702]
[WorldCat.org]
[DOI]
(I p)
Ziqing Mei, Feng Wang, Yutao Qi, Zhiyuan Zhou, Qi Hu, Han Li, Jiawei Wu, Yigong Shi
Molecular determinants of MecA as a degradation tag for the ClpCP protease.
J Biol Chem: 2009, 284(49);34366-75
[PubMed:19767395]
[WorldCat.org]
[DOI]
(I p)
Jeanette Hahn, Naomi Kramer, Kenneth Briley, David Dubnau
McsA and B mediate the delocalization of competence proteins from the cell poles of Bacillus subtilis.
Mol Microbiol: 2009, 72(1);202-15
[PubMed:19226326]
[WorldCat.org]
[DOI]
(I p)
James Kain, Gina G He, Richard Losick
Polar localization and compartmentalization of ClpP proteases during growth and sporulation in Bacillus subtilis.
J Bacteriol: 2008, 190(20);6749-57
[PubMed:18689476]
[WorldCat.org]
[DOI]
(I p)
Lyle A Simmons, Alan D Grossman, Graham C Walker
Clp and Lon proteases occupy distinct subcellular positions in Bacillus subtilis.
J Bacteriol: 2008, 190(20);6758-68
[PubMed:18689473]
[WorldCat.org]
[DOI]
(I p)
Adam Reeves, Ulf Gerth, Uwe Völker, W G Haldenwang
ClpP modulates the activity of the Bacillus subtilis stress response transcription factor, sigmaB.
J Bacteriol: 2007, 189(17);6168-75
[PubMed:17586624]
[WorldCat.org]
[DOI]
(P p)
Peter Prepiak, David Dubnau
A peptide signal for adapter protein-mediated degradation by the AAA+ protease ClpCP.
Mol Cell: 2007, 26(5);639-47
[PubMed:17560370]
[WorldCat.org]
[DOI]
(P p)
Janine Kirstein, David A Dougan, Ulf Gerth, Michael Hecker, Kürşad Turgay
The tyrosine kinase McsB is a regulated adaptor protein for ClpCP.
EMBO J: 2007, 26(8);2061-70
[PubMed:17380125]
[WorldCat.org]
[DOI]
(P p)
Stephan Zellmeier, Wolfgang Schumann, Thomas Wiegert
Involvement of Clp protease activity in modulating the Bacillus subtilissigmaw stress response.
Mol Microbiol: 2006, 61(6);1569-82
[PubMed:16899079]
[WorldCat.org]
[DOI]
(P p)
Holger Kock, Ulf Gerth, Michael Hecker
The ClpP peptidase is the major determinant of bulk protein turnover in Bacillus subtilis.
J Bacteriol: 2004, 186(17);5856-64
[PubMed:15317791]
[WorldCat.org]
[DOI]
(P p)
Holger Kock, Ulf Gerth, Michael Hecker
MurAA, catalysing the first committed step in peptidoglycan biosynthesis, is a target of Clp-dependent proteolysis in Bacillus subtilis.
Mol Microbiol: 2004, 51(4);1087-102
[PubMed:14763982]
[WorldCat.org]
[DOI]
(P p)
Ulf Gerth, Janine Kirstein, Jörg Mostertz, Torsten Waldminghaus, Marcus Miethke, Holger Kock, Michael Hecker
Fine-tuning in regulation of Clp protein content in Bacillus subtilis.
J Bacteriol: 2004, 186(1);179-91
[PubMed:14679237]
[WorldCat.org]
[DOI]
(P p)
Qi Pan, Richard Losick
Unique degradation signal for ClpCP in Bacillus subtilis.
J Bacteriol: 2003, 185(17);5275-8
[PubMed:12923101]
[WorldCat.org]
[DOI]
(P p)
Tilman Schlothauer, Axel Mogk, David A Dougan, Bernd Bukau, Kürşad Turgay
MecA, an adaptor protein necessary for ClpC chaperone activity.
Proc Natl Acad Sci U S A: 2003, 100(5);2306-11
[PubMed:12598648]
[WorldCat.org]
[DOI]
(P p)
Tiina Pummi, Soile Leskelä, Eva Wahlström, Ulf Gerth, Harold Tjalsma, Michael Hecker, Matti Sarvas, Vesa P Kontinen
ClpXP protease regulates the signal peptide cleavage of secretory preproteins in Bacillus subtilis with a mechanism distinct from that of the Ecs ABC transporter.
J Bacteriol: 2002, 184(4);1010-8
[PubMed:11807061]
[WorldCat.org]
[DOI]
(P p)
Q Pan, D A Garsin, R Losick
Self-reinforcing activation of a cell-specific transcription factor by proteolysis of an anti-sigma factor in B. subtilis.
Mol Cell: 2001, 8(4);873-83
[PubMed:11684022]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, M Brigulla, S Haas, J D Hoheisel, U Völker, M Hecker
Global analysis of the general stress response of Bacillus subtilis.
J Bacteriol: 2001, 183(19);5617-31
[PubMed:11544224]
[WorldCat.org]
[DOI]
(P p)
H Nanamiya, K Takahashi, M Fujita, F Kawamura
Deficiency of the initiation events of sporulation in Bacillus subtilis clpP mutant can be suppressed by a lack of the Spo0E protein phosphatase.
Biochem Biophys Res Commun: 2000, 279(1);229-33
[PubMed:11112444]
[WorldCat.org]
[DOI]
(P p)
E Krüger, E Witt, S Ohlmeier, R Hanschke, M Hecker
The clp proteases of Bacillus subtilis are directly involved in degradation of misfolded proteins.
J Bacteriol: 2000, 182(11);3259-65
[PubMed:10809708]
[WorldCat.org]
[DOI]
(P p)
I Derré, G Rapoport, T Msadek
CtsR, a novel regulator of stress and heat shock response, controls clp and molecular chaperone gene expression in gram-positive bacteria.
Mol Microbiol: 1999, 31(1);117-31
[PubMed:9987115]
[WorldCat.org]
[DOI]
(P p)
K Turgay, J Hahn, J Burghoorn, D Dubnau
Competence in Bacillus subtilis is controlled by regulated proteolysis of a transcription factor.
EMBO J: 1998, 17(22);6730-8
[PubMed:9890793]
[WorldCat.org]
[DOI]
(P p)
U Gerth, E Krüger, I Derré, T Msadek, M Hecker
Stress induction of the Bacillus subtilis clpP gene encoding a homologue of the proteolytic component of the Clp protease and the involvement of ClpP and ClpX in stress tolerance.
Mol Microbiol: 1998, 28(4);787-802
[PubMed:9643546]
[WorldCat.org]
[DOI]
(P p)
T Msadek, V Dartois, F Kunst, M L Herbaud, F Denizot, G Rapoport
ClpP of Bacillus subtilis is required for competence development, motility, degradative enzyme synthesis, growth at high temperature and sporulation.
Mol Microbiol: 1998, 27(5);899-914
[PubMed:9535081]
[WorldCat.org]
[DOI]
(P p)