Difference between revisions of "XynC"
| (3 intermediate revisions by 2 users not shown) | |||
| Line 27: | Line 27: | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=xynC_1942714_1943982_-1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:xynC_expression.png|500px]] | + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=xynC_1942714_1943982_-1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:xynC_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU18150]] |
|- | |- | ||
|} | |} | ||
| Line 35: | Line 35: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/> | <br/><br/> | ||
| Line 56: | Line 52: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU18150&redirect=T BSU18150] | ||
* '''DBTBS entry:''' no entry | * '''DBTBS entry:''' no entry | ||
| Line 94: | Line 91: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU18150&redirect=T BSU18150] | ||
* '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=3GTN 3GTN] {{PubMed|19407387,21256135}} | * '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=3GTN 3GTN] {{PubMed|19407387,21256135}} | ||
| Line 110: | Line 108: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=xynC_1942714_1943982_-1 xynC] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=xynC_1942714_1943982_-1 xynC] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 141: | Line 139: | ||
<pubmed>20735481 </pubmed> | <pubmed>20735481 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | + | <pubmed>19407387, 18957862, 17028274 20817675,21256135 24271172 </pubmed> | |
| − | <pubmed>19407387, 18957862, 17028274 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Latest revision as of 13:51, 2 April 2014
- Description: endo-xylanase, preference for methylglucurono-xylan
| Gene name | xynC |
| Synonyms | ynfF |
| Essential | no |
| Product | endo-xylanase |
| Function | xylan degradation |
| Gene expression levels in SubtiExpress: xynC | |
| MW, pI | 47 kDa, 9.078 |
| Gene length, protein length | 1266 bp, 422 aa |
| Immediate neighbours | ynfE, xynD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 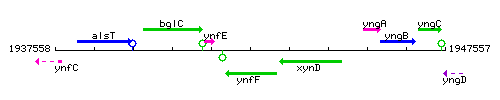
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed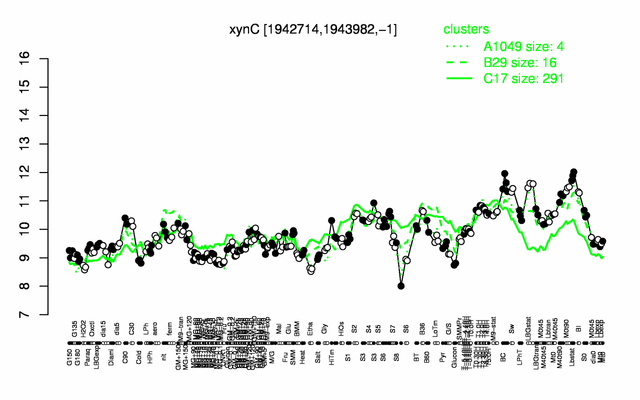
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU18150
Phenotypes of a mutant
Database entries
- BsubCyc: BSU18150
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endohydrolysis of (1->4)-beta-D-xylosyl links in some glucuronoarabinoxylans (according to Swiss-Prot)
- Protein family: glycosyl hydrolase 30 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- extracellular (signal peptide) PubMed
Database entries
- BsubCyc: BSU18150
- UniProt: Q45070
- KEGG entry: [2]
- E.C. number: 3.2.1.136
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Mun Su Rhee, Lusha Wei, Neha Sawhney, John D Rice, Franz J St John, Jason C Hurlbert, James F Preston
Engineering the xylan utilization system in Bacillus subtilis for production of acidic Xylooligosaccharides.
Appl Environ Microbiol: 2014, 80(3);917-27
[PubMed:24271172]
[WorldCat.org]
[DOI]
(I p)
Franz J St John, Jason C Hurlbert, John D Rice, James F Preston, Edwin Pozharski
Ligand bound structures of a glycosyl hydrolase family 30 glucuronoxylan xylanohydrolase.
J Mol Biol: 2011, 407(1);92-109
[PubMed:21256135]
[WorldCat.org]
[DOI]
(I p)
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
Franz J St John, David K Godwin, James F Preston, Edwin Pozharski, Jason C Hurlbert
Crystallization and crystallographic analysis of Bacillus subtilis xylanase C.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2009, 65(Pt 5);499-503
[PubMed:19407387]
[WorldCat.org]
[DOI]
(I p)
Birgit Voigt, Haike Antelmann, Dirk Albrecht, Armin Ehrenreich, Karl-Heinz Maurer, Stefan Evers, Gerhard Gottschalk, Jan Maarten van Dijl, Thomas Schweder, Michael Hecker
Cell physiology and protein secretion of Bacillus licheniformis compared to Bacillus subtilis.
J Mol Microbiol Biotechnol: 2009, 16(1-2);53-68
[PubMed:18957862]
[WorldCat.org]
[DOI]
(I p)
Franz J St John, John D Rice, James F Preston
Characterization of XynC from Bacillus subtilis subsp. subtilis strain 168 and analysis of its role in depolymerization of glucuronoxylan.
J Bacteriol: 2006, 188(24);8617-26
[PubMed:17028274]
[WorldCat.org]
[DOI]
(P p)