Difference between revisions of "YkoV"
| Line 1: | Line 1: | ||
* '''Description:''' DNA-end-binding protein Ku, recruits [[LigD]] to DNA ends, confers dry-heat resistance to dormant spores [http://www.ncbi.nlm.nih.gov/sites/entrez/16497325 PubMed] | * '''Description:''' DNA-end-binding protein Ku, recruits [[LigD]] to DNA ends, confers dry-heat resistance to dormant spores [http://www.ncbi.nlm.nih.gov/sites/entrez/16497325 PubMed] | ||
| − | + | <br/> | |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 57: | Line 57: | ||
* sensitivity to ionizing radiation in the stationary phase {{PubMed|12215643}} | * sensitivity to ionizing radiation in the stationary phase {{PubMed|12215643}} | ||
* sensitivity of spores to several DNA-damaging treatments known to cause double strand breaks, such as UV-ray, X-ray, ultrahigh vacuum and wet heat {{PubMed|16497325,17293412}} | * sensitivity of spores to several DNA-damaging treatments known to cause double strand breaks, such as UV-ray, X-ray, ultrahigh vacuum and wet heat {{PubMed|16497325,17293412}} | ||
| + | * a ''[[ykoV]]-[[ligD]]'' double mutant is sensitive to radiation {{PubMed|24123749}} | ||
| + | |||
=== Database entries === | === Database entries === | ||
| Line 82: | Line 84: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
Revision as of 16:31, 20 December 2013
- Description: DNA-end-binding protein Ku, recruits LigD to DNA ends, confers dry-heat resistance to dormant spores PubMed
| Gene name | ykoV |
| Synonyms | |
| Essential | no |
| Product | DNA-end-binding protein Ku |
| Function | non-homologous end joining DNA repair, repair of gapped DNA substrates |
| Gene expression levels in SubtiExpress: ykoV | |
| Interactions involving this protein in SubtInteract: YkoV | |
| MW, pI | 34 kDa, 7.213 |
| Gene length, protein length | 933 bp, 311 aa |
| Immediate neighbours | ligD, dgcW |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 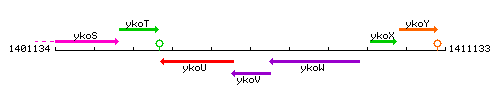
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed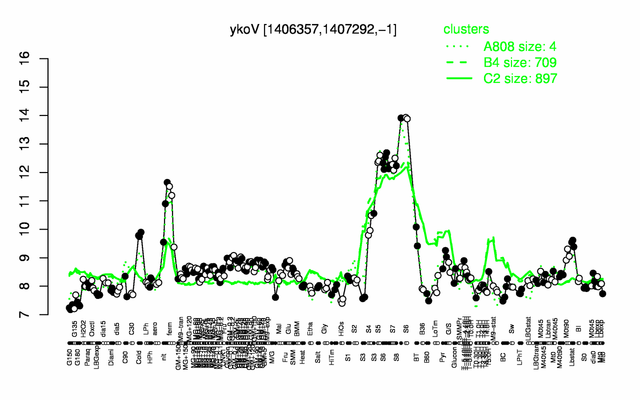
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13410
Phenotypes of a mutant
- sensitivity to ionizing radiation in the stationary phase PubMed
- sensitivity of spores to several DNA-damaging treatments known to cause double strand breaks, such as UV-ray, X-ray, ultrahigh vacuum and wet heat PubMed
- a ykoV-ligD double mutant is sensitive to radiation PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization:
- associates with the nucleoid during germination PubMed
Database entries
- Structure:
- UniProt: O34859
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- BP141 (ykoV-ligD::kan) available in Fabian Commichau's lab
- BP142 (ykoW-ykoV-ligD::kan) available in Fabian Commichau's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications