Difference between revisions of "RsiW"
| Line 22: | Line 22: | ||
|style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 624 bp, 208 aa | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 624 bp, 208 aa | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[sigW]]'', ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[sigW]]'', ''[[cdaA]]'' |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB11950]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB11950]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | ||
Revision as of 11:57, 30 November 2012
- Description: anti-SigW
| Gene name | rsiW |
| Synonyms | ybbM |
| Essential | no |
| Product | anti-SigW |
| Function | control of SigW activity |
| Gene expression levels in SubtiExpress: rsiW | |
| Interactions involving this protein in SubtInteract: RsiW | |
| MW, pI | 23 kDa, 6.646 |
| Gene length, protein length | 624 bp, 208 aa |
| Immediate neighbours | sigW, cdaA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 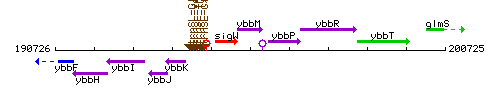
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed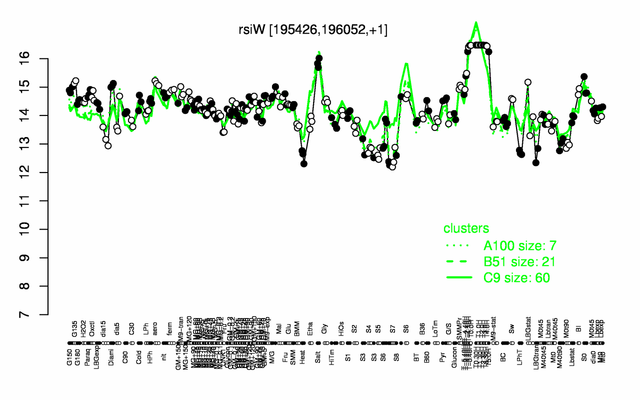
| |
Contents
Categories containing this gene/protein
sigma factors and their control, cell envelope stress proteins (controlled by SigM, V, W, X, Y), membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU01740
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: anti-sigma-W factor family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity: degraded by RasP and PrsW under conditions of alkali shock or in the presence of antimicrobial peptides, respectively. This results in the release of SigW
- Localization: membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: Q45588
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information: the mRNA is very stable (half-life > 15 min) PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Thomas Wiegert, University of Bayreuth, Germany Homepage
Your additional remarks
References
Reviews
Additional reviews: PubMed
Original Publications