Difference between revisions of "FrlD"
| Line 37: | Line 37: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | + | <br/><br/> | |
| − | |||
| − | |||
| − | |||
| − | |||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| Line 107: | Line 103: | ||
* '''Operon:''' | * '''Operon:''' | ||
** ''[[yurR]]-[[yurQ]]-[[frlB]]-[[frlO]]-[[frlN]]-[[frlM]]-[[frlD]]'' {{PubMed|12823818}} | ** ''[[yurR]]-[[yurQ]]-[[frlB]]-[[frlO]]-[[frlN]]-[[frlM]]-[[frlD]]'' {{PubMed|12823818}} | ||
| − | ** ''[[frlB]]-[[frlO]]-[[frlN]]-[[frlM]]-[[frlD]]'' {{PubMed|21347729}} | + | ** ''[[frlB]]-[[frlO]]-[[frlN]]-[[frlM]]-[[frlD]]'' {{PubMed|23175651,21347729}} |
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=frlD_3347051_3347905_-1 frlD] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=frlD_3347051_3347905_-1 frlD] {{PubMed|22383849}} | ||
| Line 123: | Line 119: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
** the mRNA is substantially stabilized upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | ** the mRNA is substantially stabilized upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | ||
| + | ** the ''[[frlB]]-[[frlO]]-[[frlN]]-[[frlM]]-[[frlD]]'' operon is not expressed in a ''[[cshA]]'' mutant {{PubMed|23175651}} | ||
=Biological materials = | =Biological materials = | ||
| Line 144: | Line 141: | ||
=References= | =References= | ||
'''Additional references:''' {{PubMed|21397556}} | '''Additional references:''' {{PubMed|21397556}} | ||
| + | <pubmed> 23175651 </pubmed> | ||
<big>''Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J'' </big> | <big>''Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J'' </big> | ||
<big>'''RNA processing in ''Bacillus subtilis'': identification of targets of the essential RNase Y.''' </big> | <big>'''RNA processing in ''Bacillus subtilis'': identification of targets of the essential RNase Y.''' </big> | ||
Revision as of 17:11, 26 November 2012
- Description: fructosamine kinase
| Gene name | frlD |
| Synonyms | yurL |
| Essential | no |
| Product | fructosamine kinase |
| Function | metabolism of sugar amines |
| Gene expression levels in SubtiExpress: frlD | |
| Metabolic function and regulation of this protein in SubtiPathways: Alternative nitrogen sources | |
| MW, pI | 30 kDa, 4.909 |
| Gene length, protein length | 852 bp, 284 aa |
| Immediate neighbours | frlR, frlM |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed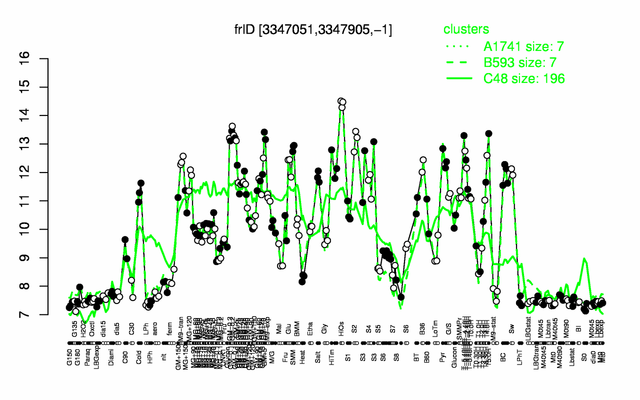
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, utilization of nitrogen sources other than amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU32570
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: carbohydrate kinase pfkB family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O32153
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional references: PubMed
Martin Lehnik-Habrink, Leonie Rempeters, Ákos T Kovács, Christoph Wrede, Claudia Baierlein, Heike Krebber, Oscar P Kuipers, Jörg Stülke
DEAD-Box RNA helicases in Bacillus subtilis have multiple functions and act independently from each other.
J Bacteriol: 2013, 195(3);534-44
[PubMed:23175651]
[WorldCat.org]
[DOI]
(I p)
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Veronika Maria Deppe, Stephanie Klatte, Johannes Bongaerts, Karl-Heinz Maurer, Timothy O'Connell, Friedhelm Meinhardt
Genetic control of amadori product degradation in Bacillus subtilis via regulation of frlBONMD expression by FrlR.
Appl Environ Microbiol: 2011, 77(9);2839-46
[PubMed:21398478]
[WorldCat.org]
[DOI]
(I p)
Veronika Maria Deppe, Johannes Bongaerts, Timothy O'Connell, Karl-Heinz Maurer, Friedhelm Meinhardt
Enzymatic deglycation of Amadori products in bacteria: mechanisms, occurrence and physiological functions.
Appl Microbiol Biotechnol: 2011, 90(2);399-406
[PubMed:21347729]
[WorldCat.org]
[DOI]
(I p)
Elsa Wiame, Pedro Lamosa, Helena Santos, Emile Van Schaftingen
Identification of glucoselysine-6-phosphate deglycase, an enzyme involved in the metabolism of the fructation product glucoselysine.
Biochem J: 2005, 392(Pt 2);263-9
[PubMed:16153181]
[WorldCat.org]
[DOI]
(I p)
Elsa Wiame, Armelle Duquenne, Ghislain Delpierre, Emile Van Schaftingen
Identification of enzymes acting on alpha-glycated amino acids in Bacillus subtilis.
FEBS Lett: 2004, 577(3);469-72
[PubMed:15556630]
[WorldCat.org]
[DOI]
(P p)
Ken-ichi Yoshida, Hirotake Yamaguchi, Masaki Kinehara, Yo-hei Ohki, Yoshiko Nakaura, Yasutaro Fujita
Identification of additional TnrA-regulated genes of Bacillus subtilis associated with a TnrA box.
Mol Microbiol: 2003, 49(1);157-65
[PubMed:12823818]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Yoshiko Nakaura, Robert P Shivers, Hirotake Yamaguchi, Richard Losick, Yasutaro Fujita, Abraham L Sonenshein
Additional targets of the Bacillus subtilis global regulator CodY identified by chromatin immunoprecipitation and genome-wide transcript analysis.
J Bacteriol: 2003, 185(6);1911-22
[PubMed:12618455]
[WorldCat.org]
[DOI]
(P p)