Difference between revisions of "SigH"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 1: | Line 1: | ||
| − | + | * '''Description:''' [[RNA polymerase]] [[sigma factor]] SigH <br/><br/> | |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || [[transcription]] of early stationary phase genes <br/>(sporulation, competence) | |style="background:#ABCDEF;" align="center"|'''Function''' || [[transcription]] of early stationary phase genes <br/>(sporulation, competence) | ||
| − | |||
| − | |||
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/SigH SigH] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/SigH SigH] | ||
| Line 28: | Line 26: | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:sigH_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:sigH_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| − | |||
| − | |||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/><br/> | <br/><br/><br/> | ||
| Line 112: | Line 102: | ||
* '''Operon:''' ''sigH'' (DBTBS) | * '''Operon:''' ''sigH'' (DBTBS) | ||
| − | * | + | * '''[[Sigma factor]]:''' [[SigA]] {{PubMed|1898930}} |
| − | |||
| − | |||
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 122: | Line 110: | ||
** [[AbrB]]: transcription repression [http://www.ncbi.nlm.nih.gov/sites/entrez/7592498 PubMed] | ** [[AbrB]]: transcription repression [http://www.ncbi.nlm.nih.gov/sites/entrez/7592498 PubMed] | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' |
=Biological materials = | =Biological materials = | ||
Revision as of 10:18, 7 August 2012
- Description: RNA polymerase sigma factor SigH
| Gene name | sigH |
| Synonyms | spo0H |
| Essential | no |
| Product | RNA polymerase sigma factor SigH |
| Function | transcription of early stationary phase genes (sporulation, competence) |
| Interactions involving this protein in SubtInteract: SigH | |
| MW, pI | 25 kDa, 5.146 |
| Gene length, protein length | 654 bp, 218 aa |
| Immediate neighbours | yacP, rpmGB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 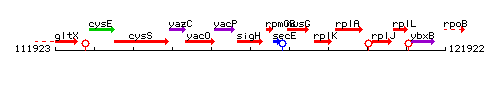
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transcription, sigma factors and their control
This gene is a member of the following regulons
The SigH regulon
The gene
Basic information
- Locus tag: BSU00980
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: sigma-70 factor family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P17869
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: sigH (DBTBS)
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Charles Moran, Emory University, NC, USA homepage
Your additional remarks
References
Reviews
Michiel J L de Hoon, Patrick Eichenberger, Dennis Vitkup
Hierarchical evolution of the bacterial sporulation network.
Curr Biol: 2010, 20(17);R735-45
[PubMed:20833318]
[WorldCat.org]
[DOI]
(I p)
The SigH regulon
Original Publications