SucC
- Description: succinyl-CoA synthetase (beta subunit)
| Gene name | sucC |
| Synonyms | |
| Essential | no |
| Product | succinyl-CoA synthetase (beta subunit) |
| Function | TCA cycle |
| Gene expression levels in SubtiExpress: sucC | |
| Interactions involving this protein in SubtInteract: SucC | |
| Metabolic function and regulation of this protein in SubtiPathways: sucC | |
| MW, pI | 41 kDa, 4.846 |
| Gene length, protein length | 1155 bp, 385 aa |
| Immediate neighbours | ylqH, sucD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 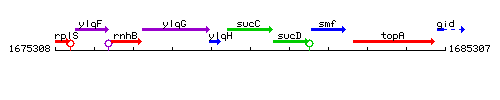
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed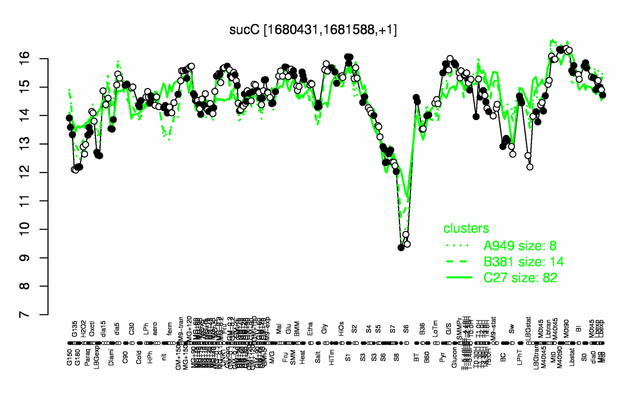
| |
Contents
Categories containing this gene/protein
ATP synthesis, carbon core metabolism, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU16090
Phenotypes of a mutant
Database entries
- BsubCyc: BSU16090
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + succinate + CoA = ADP + phosphate + succinyl-CoA (according to Swiss-Prot)
- Protein family: ATP-grasp domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: Reversible Michaelis-Menten FEBS Letters
- Modification: phosphorylation on Ser-220 PubMed
- Effectors of protein activity:
- Inhibited by 2-oxoglutarate, ATP and NADH FEBS Letters
- GTP is not accept by the enzyme FEBS Letters
- Inhibited by 2-oxoglutarate, ATP and NADH FEBS Letters
Database entries
- BsubCyc: BSU16090
- Structure: 1JKJ (E. coli)
- UniProt: P80886
- KEGG entry: [3]
- E.C. number: 6.2.1.5
Additional information
- extensive information on the structure and enzymatic properties of succinyl-CoA synthetase can be found at Proteopedia
Expression and regulation
- Regulatory mechanism:
- Additional information:
- the mRNA base-pairs with the RoxS sRNA PubMed
- the roxS-sucC RNA duplex forms a cleavage site for RNase III PubMed
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 7352 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 44785 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 10243 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 5100 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 8473 PubMed
Biological materials
- Mutant:
- 1A1006 ( sucC::spec), available at BGSC
- GP1134 (cat), available in Jörg Stülke's lab
- GP791 (sucC-sucD::tet), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References