SwrAA/1
- Description: control of DegU activity, enhances sigD transcription, controls the number of flagellar basal bodies, inactive pseudogene in strain 168
| Gene name | swrAA/1 |
| Synonyms | yvzD, swrAA, ifm |
| Essential | no |
| Product | swarming motility protein |
| Function | control of DegU activity |
| Gene expression levels in SubtiExpress: swrAA/1 | |
| Interactions involving this protein in SubtInteract: SwrA | |
| MW, pI | - , - |
| Gene length, protein length | 336 bp, - |
| Immediate neighbours | minJ, swrAA/2 |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 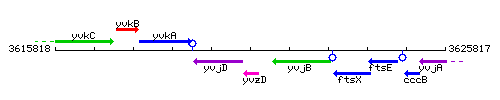
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed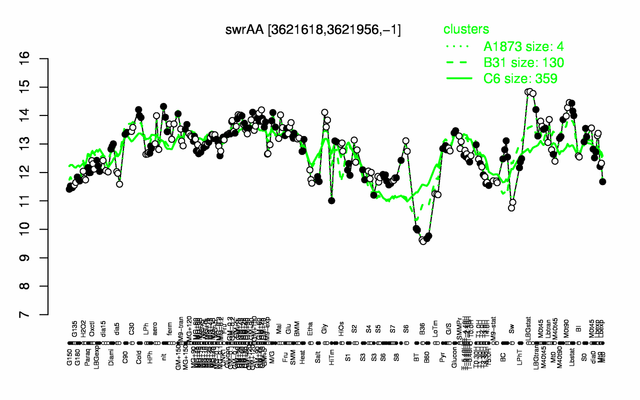
| |
Contents
Categories containing this gene/protein
biofilm formation, motility and chemotaxis, transcription factors and their control, pseudogenes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU35230
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
- This protein is functional in undomesticated strains of B.subtilis but not in laboratory strains, such as 168, because of a frameshift mutation. Therefore laboratory strains of B.subtilis are unable to swarm.
- The frameshift in strain 168 is caused by a single insertion of an adenine in the codon for Tyr-12 which leads to the premature truncation of the protein in residue 13. In addition, the C-terminal section of swrAA was predicted to be an ORF (yvzD) by the genome project.
- Correction of sfp, epsC, swrAA, and degQ as well as introduction of rapP from a plasmid present in NCIB3610 results in biofilm formation in B. subtilis 168 PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: O32266
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Additional publications: PubMed
ÜÜÜÜ
ÖÖÖ