Pgm
Revision as of 12:58, 13 January 2009 by 134.76.70.252 (talk)
- Description: phosphoglycerate mutase, glycolytic/ gluconeogenic enzyme
| Gene name | pgm |
| Synonyms | gpmI |
| Essential | yes |
| Product | 2,3-bisphosphoglycerate-independent phosphoglycerate mutase |
| Function | enzyme in glycolysis/ gluconeogenesis |
| MW, pI | 56,1 kDa, 5.21 |
| Gene length, protein length | 1533 bp, 511 amino acids |
| Immediate neighbours | tpi, eno |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 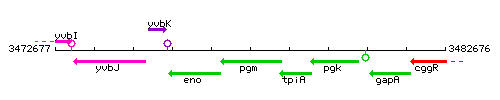
| |
Contents
The gene
Basic information
- Coordinates: 3476911 - 3478443
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2-phospho-D-glycerate = 3-phospho-D-glycerate
- Protein family: BPG-independent phosphoglycerate mutase family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation PubMed
- Cofactor(s): 2 manganese ions per subunit
- Effectors of protein activity:
- Interactions: Pgm-PfkA
- Localization: cytoplasm PubMed
Database entries
- Structure: Geobacillus stearothermophilus, complex with 2-phosphoglycerate NCBI, Geobacillus stearothermophilus, complex with 3-phosphoglycerate NCBI
- Swiss prot entry: [3]
- KEGG entry: [4]
- E.C. number: [5]
Additional information
is pH sensitive
Expression and regulation
- Sigma factor: SigA
- Additional information:
Biological materials
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
- Ludwig, H., Homuth, G., Schmalisch, M., Dyka, F. M., Hecker, M., and Stülke, J. (2001) Transcription of glycolytic genes and operons in Bacillus subtilis: evidence for the presence of multiple levels of control of the gapA operon. Mol Microbiol 41, 409-422.PubMed