MhqE
- Description: dioxygenase/glyoxalase
| Gene name | mhqE |
| Synonyms | yodE |
| Essential | no |
| Product | dioxygenase/glyoxalase |
| Function | resistance to methyl hydroquinone and catechol |
| MW, pI | 33 kDa, 5.041 |
| Gene length, protein length | 909 bp, 303 aa |
| Immediate neighbours | mhqD, yodF |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 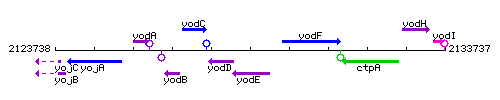
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU19570
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: O34543
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Stefanie Töwe, Montira Leelakriangsak, Kazuo Kobayashi, Nguyen Van Duy, Michael Hecker, Peter Zuber, Haike Antelmann
The MarR-type repressor MhqR (YkvE) regulates multiple dioxygenases/glyoxalases and an azoreductase which confer resistance to 2-methylhydroquinone and catechol in Bacillus subtilis.
Mol Microbiol: 2007, 66(1);40-54
[PubMed:17725564]
[WorldCat.org]
[DOI]
(P p)
Van Duy Nguyen, Carmen Wolf, Ulrike Mäder, Michael Lalk, Peter Langer, Ulrike Lindequist, Michael Hecker, Haike Antelmann
Transcriptome and proteome analyses in response to 2-methylhydroquinone and 6-brom-2-vinyl-chroman-4-on reveal different degradation systems involved in the catabolism of aromatic compounds in Bacillus subtilis.
Proteomics: 2007, 7(9);1391-408
[PubMed:17407181]
[WorldCat.org]
[DOI]
(P p)
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed