Difference between revisions of "LeuB"
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || biosynthesis of leucine | |style="background:#ABCDEF;" align="center"|'''Function''' || biosynthesis of leucine | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/CoA_synthesis.html Coenzyme A]''' | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/ile_val_leu.html Ile, Leu, Val], [http://subtiwiki.uni-goettingen.de/pathways/CoA_synthesis.html Coenzyme A]''' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 39 kDa, 4.744 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 39 kDa, 4.744 | ||
Revision as of 14:05, 17 June 2009
- Description: 3-isopropylmalate dehydrogenase
| Gene name | leuB |
| Synonyms | leuC |
| Essential | no |
| Product | 3-isopropylmalate dehydrogenase |
| Function | biosynthesis of leucine |
| Metabolic function and regulation of this protein in SubtiPathways: Ile, Leu, Val, Coenzyme A | |
| MW, pI | 39 kDa, 4.744 |
| Gene length, protein length | 1095 bp, 365 aa |
| Immediate neighbours | leuC, leuA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 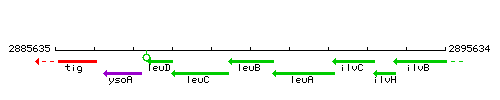
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU28270
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (2R,3S)-3-isopropylmalate + NAD+ = 4-methyl-2-oxopentanoate + CO2 + NADH (according to Swiss-Prot)
- Protein family: LeuB type 1 subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure: 1XAD (Thermus thermophilus)
- Swiss prot entry: P05645
- KEGG entry: [3]
- E.C. number: 1.1.1.85
Additional information
Expression and regulation
- Regulation: repressed by casamino acids PubMed , expressed in the absence of branched-chain amino acids (BCAA), expression is stimulated in the presence of glucose PubMed, repressed by CodY PubMed
- Regulatory mechanism: CodY: transcription repression PubMed, glucose regulation: CcpA PubMed, repression by BCAA: tRNA-controlled RNA switch (T-box) that mediates termination/antitermination
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References