Difference between revisions of "LysC"
(→Other original Publications) |
(→Other original Publications) |
||
| Line 153: | Line 153: | ||
===Other original Publications=== | ===Other original Publications=== | ||
| − | <pubmed>2168395, 1624109, 2559145, 2557260,12850135 12107147, 17981983 15378759 25935345</pubmed> | + | <pubmed>2168395, 1624109, 2559145, 2557260,12850135 12107147, 17981983 15378759 25935345 2153658 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 15:31, 19 May 2015
- Description: aspartokinase II (alpha and beta subunits)
| Gene name | lysC |
| Synonyms | ask, aecA |
| Essential | no |
| Product | aspartokinase II (alpha and beta subunits) |
| Function | biosynthesis of lysine |
| Gene expression levels in SubtiExpress: lysC | |
| Metabolic function and regulation of this protein in SubtiPathways: lysC | |
| MW, pI | 43 kDa, 4.643 |
| Gene length, protein length | 1224 bp, 408 aa |
| Immediate neighbours | yslB, uvrC |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 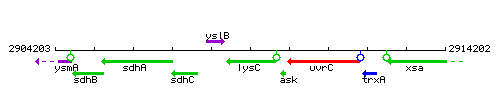
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed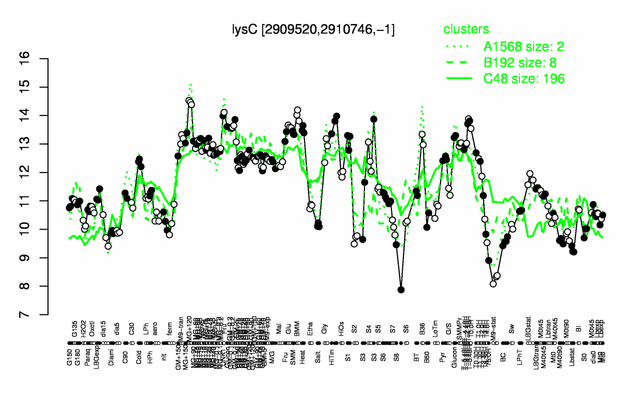
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28470
Phenotypes of a mutant
Database entries
- BsubCyc: BSU28470
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + L-aspartate = ADP + 4-phospho-L-aspartate (according to Swiss-Prot)
- Protein family: aspartokinase family (according to Swiss-Prot)
- Paralogous protein(s): DapG
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- BsubCyc: BSU28470
- Structure: 2RE1 (from Neisseria meningitidis mc58, 40% identity, 58% similarity)
- UniProt: P08495
- KEGG entry: [3]
- E.C. number: 2.7.2.4
Additional information
Expression and regulation
- Operon: lysC PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
- subject to Clp-dependent proteolysis upon glucose starvation PubMed, also degraded upon ammonium or amino acid starvation PubMed
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 1459 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 4406 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 1282 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 806 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original Publications
The L-box riboswitch
Other original Publications
Junming Wang, Dongfang Gao, Xiaoli Yu, Wen Li, Qingsheng Qi
Evolution of a chimeric aspartate kinase for L-lysine production using a synthetic RNA device.
Appl Microbiol Biotechnol: 2015, 99(20);8527-36
[PubMed:25935345]
[WorldCat.org]
[DOI]
(I p)
Ulf Gerth, Holger Kock, Ilja Kusters, Stephan Michalik, Robert L Switzer, Michael Hecker
Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis.
J Bacteriol: 2008, 190(1);321-31
[PubMed:17981983]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)
Y Lu, T N Shevtchenko, H Paulus
Fine-structure mapping of cis-acting control sites in the lysC operon of Bacillus subtilis.
FEMS Microbiol Lett: 1992, 71(1);23-7
[PubMed:1624109]
[WorldCat.org]
[DOI]
(P p)
L M Graves, R L Switzer
Aspartokinase II from Bacillus subtilis is degraded in response to nutrient limitation.
J Biol Chem: 1990, 265(25);14947-55
[PubMed:2168395]
[WorldCat.org]
(P p)
J J Zhang, F M Hu, N Y Chen, H Paulus
Comparison of the three aspartokinase isozymes in Bacillus subtilis Marburg and 168.
J Bacteriol: 1990, 172(2);701-8
[PubMed:2153658]
[WorldCat.org]
[DOI]
(P p)
N Y Chen, J J Zhang, H Paulus
Chromosomal location of the Bacillus subtilis aspartokinase II gene and nucleotide sequence of the adjacent genes homologous to uvrC and trx of Escherichia coli.
J Gen Microbiol: 1989, 135(11);2931-40
[PubMed:2559145]
[WorldCat.org]
[DOI]
(P p)
M Petricek, L Rutberg, L Hederstedt
The structural gene for aspartokinase II in Bacillus subtilis is closely linked to the sdh operon.
FEMS Microbiol Lett: 1989, 52(1-2);85-7
[PubMed:2557260]
[WorldCat.org]
[DOI]
(P p)