Difference between revisions of "QueE"
| Line 15: | Line 15: | ||
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13740 queE] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13740 queE] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=QueE QueE] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 26 kDa, 4.962 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 26 kDa, 4.962 | ||
| Line 84: | Line 86: | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
| + | ** [[QueE]]-[[YkuN]] (electron transfer) {{PubMed|25933252}} | ||
* '''[[Localization]]:''' | * '''[[Localization]]:''' | ||
| Line 135: | Line 138: | ||
=References= | =References= | ||
| − | <pubmed>14660578,19285444, 19354300 17384645 24362703 23194065</pubmed> | + | <pubmed>14660578,19285444, 19354300 17384645 24362703 23194065 25933252</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Latest revision as of 09:31, 4 May 2015
- Description: 7-carboxy-7-deazaguanine (CDG) synthase, required for the synthesis of the modified ribonucleotide queuosine
| Gene name | queE |
| Synonyms | ykvL |
| Essential | no |
| Product | 7-carboxy-7-deazaguanine (CDG) synthase |
| Function | tRNA modification |
| Gene expression levels in SubtiExpress: queE | |
| Interactions involving this protein in SubtInteract: QueE | |
| MW, pI | 26 kDa, 4.962 |
| Gene length, protein length | 729 bp, 243 aa |
| Immediate neighbours | queD, queF |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 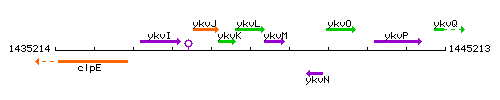
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed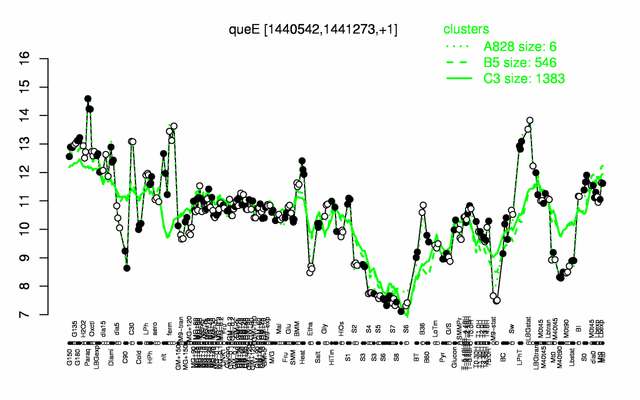
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13740
Phenotypes of a mutant
Database entries
- BsubCyc: BSU13740
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- conversion of 6-carboxy-5,6,7,8-tetrahydropterin to 7-carboxy-7-deazaguanine (CDG) PubMed
- Catalyzed reaction/ biological activity:
- Protein family: radical S-adenosyl-L-methionine (SAM) superfamily
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): [4Fe-4S] cluster, S-adenosyl-L-methionine, Mg(2+) PubMed
- Effectors of protein activity:
Database entries
- BsubCyc: BSU13740
- Structure:
- UniProt: O31677
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- repressed in the presence of queuosine (preQ1 riboswitch) PubMed
- Regulatory mechanism:
- preQ1 riboswitch: transcriptional antitermination in the absence of queuosine PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Nathan A Bruender, Anthony P Young, Vahe Bandarian
Chemical and Biological Reduction of the Radical SAM Enzyme 7-Carboxy-7-deazaguanine [corrected] Synthase.
Biochemistry: 2015, 54(18);2903-10
[PubMed:25933252]
[WorldCat.org]
[DOI]
(I p)
Daniel P Dowling, Nathan A Bruender, Anthony P Young, Reid M McCarty, Vahe Bandarian, Catherine L Drennan
Radical SAM enzyme QueE defines a new minimal core fold and metal-dependent mechanism.
Nat Chem Biol: 2014, 10(2);106-12
[PubMed:24362703]
[WorldCat.org]
[DOI]
(I p)
Reid M McCarty, Carsten Krebs, Vahe Bandarian
Spectroscopic, steady-state kinetic, and mechanistic characterization of the radical SAM enzyme QueE, which catalyzes a complex cyclization reaction in the biosynthesis of 7-deazapurines.
Biochemistry: 2013, 52(1);188-98
[PubMed:23194065]
[WorldCat.org]
[DOI]
(I p)
Reid M McCarty, Arpád Somogyi, Guangxin Lin, Neil E Jacobsen, Vahe Bandarian
The deazapurine biosynthetic pathway revealed: in vitro enzymatic synthesis of PreQ(0) from guanosine 5'-triphosphate in four steps.
Biochemistry: 2009, 48(18);3847-52
[PubMed:19354300]
[WorldCat.org]
[DOI]
(I p)
Mijeong Kang, Robert Peterson, Juli Feigon
Structural Insights into riboswitch control of the biosynthesis of queuosine, a modified nucleotide found in the anticodon of tRNA.
Mol Cell: 2009, 33(6);784-90
[PubMed:19285444]
[WorldCat.org]
[DOI]
(I p)
Adam Roth, Wade C Winkler, Elizabeth E Regulski, Bobby W K Lee, Jinsoo Lim, Inbal Jona, Jeffrey E Barrick, Ankita Ritwik, Jane N Kim, Rüdiger Welz, Dirk Iwata-Reuyl, Ronald R Breaker
A riboswitch selective for the queuosine precursor preQ1 contains an unusually small aptamer domain.
Nat Struct Mol Biol: 2007, 14(4);308-17
[PubMed:17384645]
[WorldCat.org]
[DOI]
(P p)
John S Reader, David Metzgar, Paul Schimmel, Valérie de Crécy-Lagard
Identification of four genes necessary for biosynthesis of the modified nucleoside queuosine.
J Biol Chem: 2004, 279(8);6280-5
[PubMed:14660578]
[WorldCat.org]
[DOI]
(P p)