Difference between revisions of "Rnz"
| Line 120: | Line 120: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
** number of protein molecules per cell (minimal medium with glucose and ammonium): 55 {{PubMed|24696501}} | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 55 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 278 {{PubMed|21395229}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 379 {{PubMed|21395229}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 311 {{PubMed|21395229}} | ||
=Biological materials = | =Biological materials = | ||
| − | |||
* '''Mutant:''' | * '''Mutant:''' | ||
Revision as of 14:20, 17 April 2014
- Description: RNase Z
| Gene name | rnz |
| Synonyms | yqjK |
| Essential | yes PubMed |
| Product | endoribonuclease Z |
| Function | processing of CCA-less tRNA precursors |
| Gene expression levels in SubtiExpress: rnz | |
| MW, pI | 33 kDa, 5.805 |
| Gene length, protein length | 921 bp, 307 aa |
| Immediate neighbours | yqjL, zwf |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 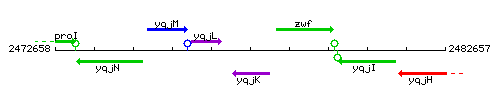
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed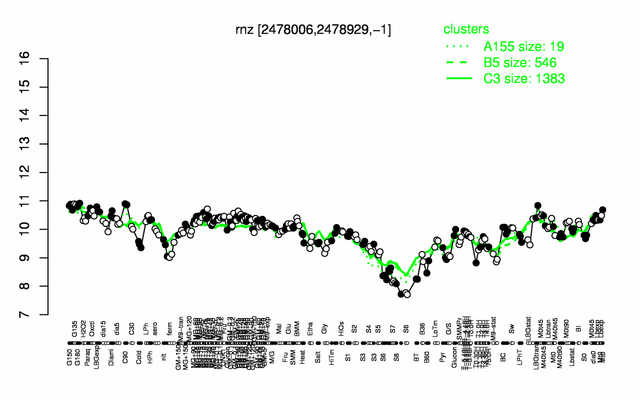
| |
Contents
Categories containing this gene/protein
Rnases, translation, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23840
Phenotypes of a mutant
essential PubMed
Database entries
- BsubCyc: BSU23840
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endonucleolytic cleavage of RNA, removing extra 3' nucleotides from tRNA precursor, generating 3' termini of tRNAs (according to Swiss-Prot)
- Protein family: RNase Z family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU23840
- UniProt: P54548
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: rnz (according to DBTBS)
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 55 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 278 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 379 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 311 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Ciaran Condon, IBPC, Paris, France Homepage
Your additional remarks
References
Reviews
Original Publications