Difference between revisions of "HisC"
| Line 121: | Line 121: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
** the mRNA is substantially stabilized upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | ** the mRNA is substantially stabilized upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 2715 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 2096 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 09:26, 17 April 2014
- Description: histidinol-phosphate aminotransferase / tyrosine and phenylalanine aminotransferase
| Gene name | hisC |
| Synonyms | aroJ |
| Essential | no |
| Product | histidinol-phosphate aminotransferase / tyrosine and phenylalanine aminotransferase |
| Function | biosynthesis of aromatic amino acids |
| Gene expression levels in SubtiExpress: hisC | |
| Metabolic function and regulation of this protein in SubtiPathways: hisC | |
| MW, pI | 39 kDa, 5.005 |
| Gene length, protein length | 1080 bp, 360 aa |
| Immediate neighbours | tyrA, trpA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 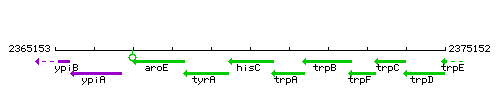
| |
Expression at a glance PubMed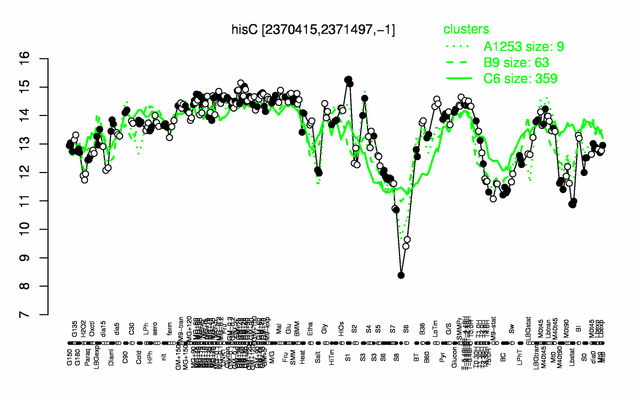
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22620
Phenotypes of a mutant
Database entries
- BsubCyc: BSU22620
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-histidinol phosphate + 2-oxoglutarate = 3-(imidazol-4-yl)-2-oxopropyl phosphate + L-glutamate (according to Swiss-Prot)
- Protein family: bacterial solute-binding protein 3 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU22620
- Swiss prot entry: P17731
- KEGG entry: [3]
- E.C. number: 2.6.1.9
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Martin Bommer, John M Ward
A 1-step microplate method for assessing the substrate range of l-α-amino acid aminotransferase.
Enzyme Microb Technol: 2013, 52(4-5);218-25
[PubMed:23540922]
[WorldCat.org]
[DOI]
(I p)
Martin Lehnik-Habrink, Marc Schaffer, Ulrike Mäder, Christine Diethmaier, Christina Herzberg, Jörg Stülke
RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y.
Mol Microbiol: 2011, 81(6);1459-73
[PubMed:21815947]
[WorldCat.org]
[DOI]
(I p)
J Sivaraman, Y Li, R Larocque, J D Schrag, M Cygler, A Matte
Crystal structure of histidinol phosphate aminotransferase (HisC) from Escherichia coli, and its covalent complex with pyridoxal-5'-phosphate and l-histidinol phosphate.
J Mol Biol: 2001, 311(4);761-76
[PubMed:11518529]
[WorldCat.org]
[DOI]
(P p)
J Otridge, P Gollnick
MtrB from Bacillus subtilis binds specifically to trp leader RNA in a tryptophan-dependent manner.
Proc Natl Acad Sci U S A: 1993, 90(1);128-32
[PubMed:8419914]
[WorldCat.org]
[DOI]
(P p)
P Babitzke, P Gollnick, C Yanofsky
The mtrAB operon of Bacillus subtilis encodes GTP cyclohydrolase I (MtrA), an enzyme involved in folic acid biosynthesis, and MtrB, a regulator of tryptophan biosynthesis.
J Bacteriol: 1992, 174(7);2059-64
[PubMed:1551827]
[WorldCat.org]
[DOI]
(P p)
D J Henner, L Band, H Shimotsu
Nucleotide sequence of the Bacillus subtilis tryptophan operon.
Gene: 1985, 34(2-3);169-77
[PubMed:3924737]
[WorldCat.org]
[DOI]
(P p)
H Shimotsu, D J Henner
Characterization of the Bacillus subtilis tryptophan promoter region.
Proc Natl Acad Sci U S A: 1984, 81(20);6315-9
[PubMed:6436812]
[WorldCat.org]
[DOI]
(P p)
D A Weigent, E W Nester
Regulation of histidinol phosphate aminotransferase synthesis by tryptophan in Bacillus subtilis.
J Bacteriol: 1976, 128(1);202-11
[PubMed:824269]
[WorldCat.org]
[DOI]
(P p)
E W Nester, A L Montoya
An enzyme common to histidine and aromatic amino acid biosynthesis in Bacillus subtilis.
J Bacteriol: 1976, 126(2);699-705
[PubMed:4431]
[WorldCat.org]
[DOI]
(P p)