Difference between revisions of "PurL"
| Line 58: | Line 58: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU06480&redirect=T BSU06480] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/purEKBCSQLFMNHD.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/purEKBCSQLFMNHD.html] | ||
| Line 93: | Line 94: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU06480&redirect=T BSU06480] | ||
* '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=3D54 3D54] (from ''Thermotoga maritima'', 39% identity, 58% similarity) {{PubMed|18597481}} | * '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=3D54 3D54] (from ''Thermotoga maritima'', 39% identity, 58% similarity) {{PubMed|18597481}} | ||
Revision as of 13:08, 2 April 2014
- Description: phosphoribosylformylglycinamidine synthase
| Gene name | purL |
| Synonyms | |
| Essential | no |
| Product | phosphoribosylformylglycinamidine synthase |
| Function | purine biosynthesis |
| Gene expression levels in SubtiExpress: purL | |
| Interactions involving this protein in SubtInteract: PurL | |
| Metabolic function and regulation of this protein in SubtiPathways: purL | |
| MW, pI | 80 kDa, 4.554 |
| Gene length, protein length | 2226 bp, 742 aa |
| Immediate neighbours | purQ, purF |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 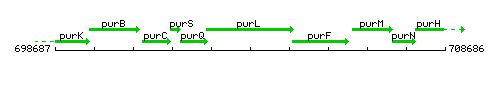
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed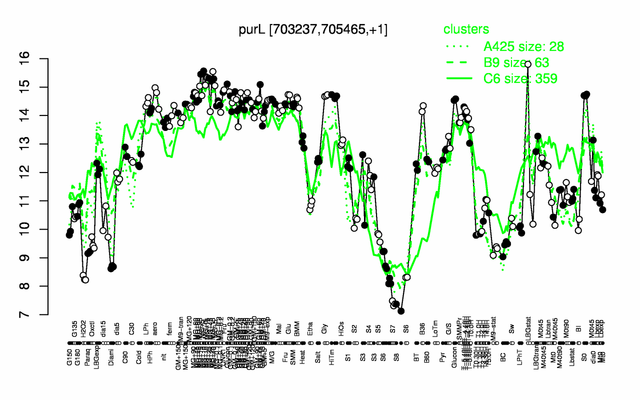
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of nucleotides, membrane proteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU06480
Phenotypes of a mutant
Database entries
- BsubCyc: BSU06480
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + N(2)-formyl-N(1)-(5-phospho-D-ribosyl)glycinamide + L-glutamine + H2O = ADP + phosphate + 2-(formamido)-N(1)-(5-phospho-D-ribosyl)acetamidine + L-glutamate (according to Swiss-Prot)
- Protein family: FGAMS family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot), membrane associated PubMed
Database entries
- BsubCyc: BSU06480
- UniProt: P12042
- KEGG entry: [3]
- E.C. number: 6.3.5.3
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
- belongs to the 100 most abundant proteins PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Yanfei Xia, Shanshan Xie, Xin Ma, Huijun Wu, Xuan Wang, Xuewen Gao
The purL gene of Bacillus subtilis is associated with nematicidal activity.
FEMS Microbiol Lett: 2011, 322(2);99-107
[PubMed:21671997]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
Aaron A Hoskins, Ruchi Anand, Steven E Ealick, JoAnne Stubbe
The formylglycinamide ribonucleotide amidotransferase complex from Bacillus subtilis: metabolite-mediated complex formation.
Biochemistry: 2004, 43(32);10314-27
[PubMed:15301530]
[WorldCat.org]
[DOI]
(P p)
Lars Engholm Johansen, Per Nygaard, Catharina Lassen, Yvonne Agersø, Hans H Saxild
Definition of a second Bacillus subtilis pur regulon comprising the pur and xpt-pbuX operons plus pbuG, nupG (yxjA), and pbuE (ydhL).
J Bacteriol: 2003, 185(17);5200-9
[PubMed:12923093]
[WorldCat.org]
[DOI]
(P p)
M Weng, P L Nagy, H Zalkin
Identification of the Bacillus subtilis pur operon repressor.
Proc Natl Acad Sci U S A: 1995, 92(16);7455-9
[PubMed:7638212]
[WorldCat.org]
[DOI]
(P p)
D J Ebbole, H Zalkin
Cloning and characterization of a 12-gene cluster from Bacillus subtilis encoding nine enzymes for de novo purine nucleotide synthesis.
J Biol Chem: 1987, 262(17);8274-87
[PubMed:3036807]
[WorldCat.org]
(P p)