Difference between revisions of "SucD"
| Line 39: | Line 39: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | + | <br/><br/> | |
| − | |||
| − | |||
| − | |||
| − | |||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
{{SubtiWiki category|[[ATP synthesis]]}}, | {{SubtiWiki category|[[ATP synthesis]]}}, | ||
{{SubtiWiki category|[[carbon core metabolism]]}}, | {{SubtiWiki category|[[carbon core metabolism]]}}, | ||
| − | {{SubtiWiki category|[[phosphoproteins]]}} | + | {{SubtiWiki category|[[phosphoproteins]]}}, |
| + | [[most abundant proteins]] | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
| Line 68: | Line 65: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 86: | Line 80: | ||
* '''Kinetic information:''' Reversible Michaelis-Menten [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6T36-44C8RWC-SH&_user=5731894&_coverDate=01%2F01%2F1985&_rdoc=28&_fmt=high&_orig=browse&_srch=doc-info(%23toc%234938%231985%23998209998%23270526%23FLP%23display%23Volume)&_cdi=4938&_sort=d&_docanchor=&_ct=40&_acct=C000043105&_version=1&_urlVersion=0&_userid=5731894&md5=f14f4734123ab1177d7217cab6c7ce7d FEBS Letters] | * '''Kinetic information:''' Reversible Michaelis-Menten [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6T36-44C8RWC-SH&_user=5731894&_coverDate=01%2F01%2F1985&_rdoc=28&_fmt=high&_orig=browse&_srch=doc-info(%23toc%234938%231985%23998209998%23270526%23FLP%23display%23Volume)&_cdi=4938&_sort=d&_docanchor=&_ct=40&_acct=C000043105&_version=1&_urlVersion=0&_userid=5731894&md5=f14f4734123ab1177d7217cab6c7ce7d FEBS Letters] | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' phosphorylation on (Ser-19 OR Thr-20) [http://www.ncbi.nlm.nih.gov/sites/entrez/17218307 PubMed] | * '''Modification:''' phosphorylation on (Ser-19 OR Thr-20) [http://www.ncbi.nlm.nih.gov/sites/entrez/17218307 PubMed] | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 120: | Line 114: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=sucD_1681617_1682519_1 sucD] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=sucD_1681617_1682519_1 sucD] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
* '''Regulation:''' repressed by glucose (2.4-fold) ([[CcpA]]) {{PubMed|12850135}} | * '''Regulation:''' repressed by glucose (2.4-fold) ([[CcpA]]) {{PubMed|12850135}} | ||
| Line 127: | Line 121: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| + | ** belongs to the 100 [[most abundant proteins]] {{PubMed|15378759}} | ||
=Biological materials = | =Biological materials = | ||
| Line 153: | Line 148: | ||
=References= | =References= | ||
| − | <pubmed>12850135 17218307 11976317 20933603</pubmed> | + | <pubmed>12850135 17218307 11976317 20933603 15378759</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 16:15, 5 March 2014
- Description: succinyl-CoA synthetase (alpha subunit)
| Gene name | sucD |
| Synonyms | |
| Essential | no |
| Product | succinyl-CoA synthetase (alpha subunit) |
| Function | TCA cycle |
| Gene expression levels in SubtiExpress: sucD | |
| Interactions involving this protein in SubtInteract: SucD | |
| Metabolic function and regulation of this protein in SubtiPathways: sucD | |
| MW, pI | 31 kDa, 5.587 |
| Gene length, protein length | 900 bp, 300 aa |
| Immediate neighbours | sucC, dprA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 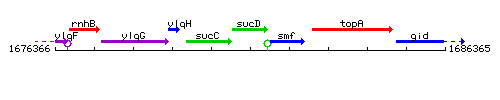
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed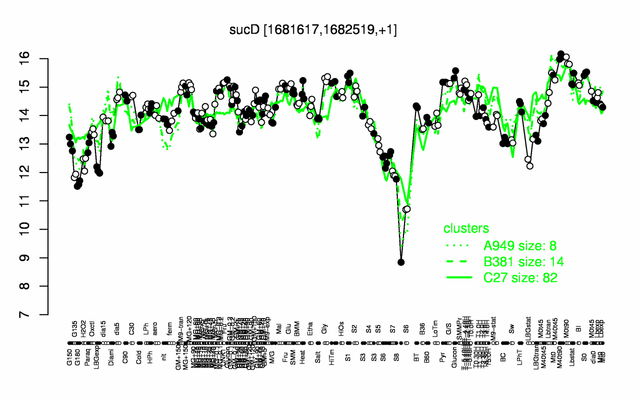
| |
Contents
Categories containing this gene/protein
ATP synthesis, carbon core metabolism, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU16100
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + succinate + CoA = ADP + phosphate + succinyl-CoA (according to Swiss-Prot)
- Protein family: succinate/malate CoA ligase alpha subunit family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: Reversible Michaelis-Menten FEBS Letters
- Modification: phosphorylation on (Ser-19 OR Thr-20) PubMed
- Effectors of protein activity:
- Inhibited by 2-oxoglutarate, ATP and NADH FEBS Letters
- GTP is not accepted by the enzyme FEBS Letters
Database entries
- Structure: 1JKJ (E. coli)
- UniProt: P80865
- KEGG entry: [3]
- E.C. number: 6.2.1.5
Additional information
- extensive information on the structure and enzymatic properties of succinyl-CoA synthetase can be found at Proteopedia
Expression and regulation
- Additional information:
- belongs to the 100 most abundant proteins PubMed
Biological materials
- Mutant:
- GP720 (spc), available in Jörg Stülke's lab
- 1A1008 ( sucD::spec), PubMed, available at BGSC
- GP791 (sucC-sucD::tet), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion: GP1435 (spc, based on pGP1870), available in Jörg Stülke's lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody:
- FLAG-tag construct: GP1425 (spc, based on pGP1331), available in Jörg Stülke's lab
Labs working on this gene/protein
Your additional remarks
References
Frederik M Meyer, Jan Gerwig, Elke Hammer, Christina Herzberg, Fabian M Commichau, Uwe Völker, Jörg Stülke
Physical interactions between tricarboxylic acid cycle enzymes in Bacillus subtilis: evidence for a metabolon.
Metab Eng: 2011, 13(1);18-27
[PubMed:20933603]
[WorldCat.org]
[DOI]
(I p)
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Ciarán Condon, Jordi Rourera, Dominique Brechemier-Baey, Harald Putzer
Ribonuclease M5 has few, if any, mRNA substrates in Bacillus subtilis.
J Bacteriol: 2002, 184(10);2845-9
[PubMed:11976317]
[WorldCat.org]
[DOI]
(P p)