Difference between revisions of "PurB"
| Line 36: | Line 36: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | + | <br/><br/> | |
| − | |||
| − | |||
| − | |||
| − | |||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| − | {{SubtiWiki category|[[biosynthesis/ acquisition of nucleotides]]}} | + | {{SubtiWiki category|[[biosynthesis/ acquisition of nucleotides]]}}, |
| + | [[most abundant proteins]] | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
| Line 64: | Line 61: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 82: | Line 76: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 113: | Line 107: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=purB_700232_701527_1 purB] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=purB_700232_701527_1 purB] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/3036807 PubMed] | + | * '''[[Sigma factor]]:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/3036807 PubMed] |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 125: | Line 119: | ||
** [[G-box]]: transcription termination/ antitermination ([[riboswitch]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/3036807 PubMed1] [http://www.ncbi.nlm.nih.gov/sites/entrez/12787499 PubMed2] | ** [[G-box]]: transcription termination/ antitermination ([[riboswitch]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/3036807 PubMed1] [http://www.ncbi.nlm.nih.gov/sites/entrez/12787499 PubMed2] | ||
| − | * '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | + | * '''Additional information:''' |
| + | ** subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | ||
| + | ** belongs to the 100 [[most abundant proteins]] {{PubMed|15378759}} | ||
=Biological materials = | =Biological materials = | ||
| Line 147: | Line 143: | ||
=References= | =References= | ||
| − | <pubmed>1722815,3036807,12923093,,7638212, </pubmed> | + | <pubmed>1722815,3036807,12923093,15378759,7638212, </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 14:44, 5 March 2014
- Description: adenylsuccinate lyase
| Gene name | purB |
| Synonyms | |
| Essential | no |
| Product | adenylsuccinate lyase |
| Function | purine biosynthesis |
| Gene expression levels in SubtiExpress: purB | |
| Metabolic function and regulation of this protein in SubtiPathways: purB | |
| MW, pI | 49 kDa, 5.822 |
| Gene length, protein length | 1293 bp, 431 aa |
| Immediate neighbours | purK, purC |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 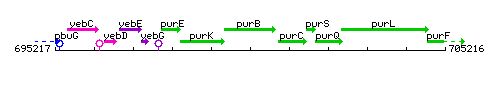
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed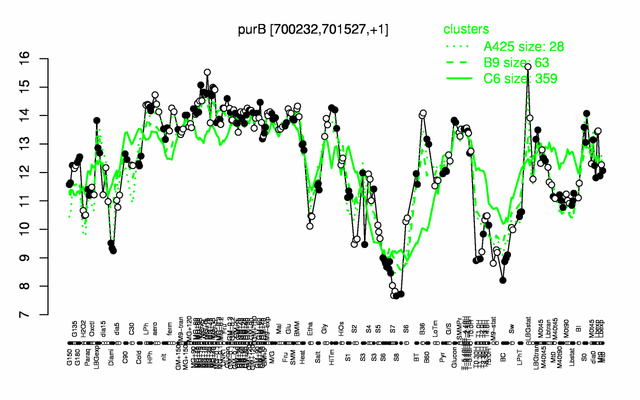
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of nucleotides, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU06440
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: N(6)-(1,2-dicarboxyethyl)AMP = fumarate + AMP (according to Swiss-Prot)
- Protein family: Adenylosuccinate lyase subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- Structure: 1F1O
- UniProt: P12047
- KEGG entry: [3]
- E.C. number: 4.3.2.2
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
- belongs to the 100 most abundant proteins PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References