Difference between revisions of "ScpA"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' part of the [[condensin]] complex, chromosomal origin condensation and segregation <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || DNA segregation and condensation protein | |style="background:#ABCDEF;" align="center"| '''Product''' || DNA segregation and condensation protein | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || segregation of replication origins |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU23220 scpA] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU23220 scpA] | ||
| Line 53: | Line 53: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | + | * essential [http://www.ncbi.nlm.nih.gov/pubmed/12682299 PubMed] | |
| − | essential [http://www.ncbi.nlm.nih.gov/pubmed/12682299 PubMed] | + | * ''[[scpA]]'' mutants are not viable on complex medim that allow rapid growth, but they are viable under conditions of slow growth {{PubMed|24440399,24440393}} |
=== Database entries === | === Database entries === | ||
| Line 79: | Line 79: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
| + | ** part of the [[condensin]] complex {{PubMed|24440399,24440393}} | ||
** present as stable monomer {{PubMed|22385855}} | ** present as stable monomer {{PubMed|22385855}} | ||
** [[ScpA]]-[[Smc]] {{PubMed|23353789,22385855,12100548}} | ** [[ScpA]]-[[Smc]] {{PubMed|23353789,22385855,12100548}} | ||
| Line 94: | Line 95: | ||
* '''[[Localization]]:''' | * '''[[Localization]]:''' | ||
** nucleoid (Multiple) [http://www.ncbi.nlm.nih.gov/sites/entrez/16479537 PubMed] | ** nucleoid (Multiple) [http://www.ncbi.nlm.nih.gov/sites/entrez/16479537 PubMed] | ||
| − | ** the ([[Smc]])2-[[ScpA]]-[[ScpB]] complex forms two bipolar assemblies on the chromosome, one in each cell half {{PubMed|23475963}} | + | ** the [[condensin]] ([[Smc]])2-[[ScpA]]-[[ScpB]] complex forms two bipolar assemblies on the chromosome, one in each cell half, localization depends on [[Spo0J]] {{PubMed|24440393,23475963}} |
=== Database entries === | === Database entries === | ||
| Line 148: | Line 149: | ||
== Original publications == | == Original publications == | ||
| − | <pubmed>12100548,12421306,12065423,7934830, 16479537, 19450516 15009890 11948165 23353789 23475963 22385855</pubmed> | + | <pubmed>12100548,12421306,12065423,7934830, 16479537, 19450516 15009890 11948165 23353789 23475963 22385855 24440399,24440393</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 15:28, 16 February 2014
- Description: part of the condensin complex, chromosomal origin condensation and segregation
| Gene name | scpA |
| Synonyms | ypuG |
| Essential | yes PubMed |
| Product | DNA segregation and condensation protein |
| Function | segregation of replication origins |
| Gene expression levels in SubtiExpress: scpA | |
| Interactions involving this protein in SubtInteract: ScpA | |
| MW, pI | 29 kDa, 4.788 |
| Gene length, protein length | 753 bp, 251 aa |
| Immediate neighbours | scpB, ypuF |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 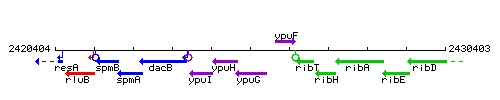
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed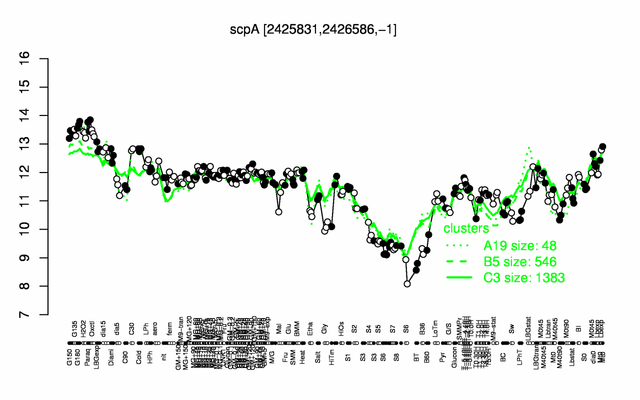
| |
Contents
Categories containing this gene/protein
DNA condensation/ segregation, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23220
Phenotypes of a mutant
- essential PubMed
- scpA mutants are not viable on complex medim that allow rapid growth, but they are viable under conditions of slow growth PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: scpA family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P35154
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications