Difference between revisions of "Smc"
| Line 30: | Line 30: | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:smc_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:smc_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=smc_1666560_1670120_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:smc_expression.png|500px]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | |||
<br/><br/> | <br/><br/> | ||
Revision as of 09:27, 19 April 2012
- Description: chromosome condensation and segregation SMC protein
| Gene name | smc |
| Synonyms | ylqA |
| Essential | yes PubMed |
| Product | chromosome condensation and segregation SMC protein |
| Function | structural maintenance of the chromosome |
| Interactions involving this protein in SubtInteract: Smc | |
| Metabolic function and regulation of this protein in SubtiPathways: Protein secretion | |
| MW, pI | 135 kDa, 5.266 |
| Gene length, protein length | 3558 bp, 1186 aa |
| Immediate neighbours | rnc, ftsY |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 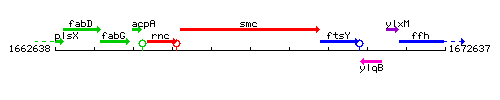
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed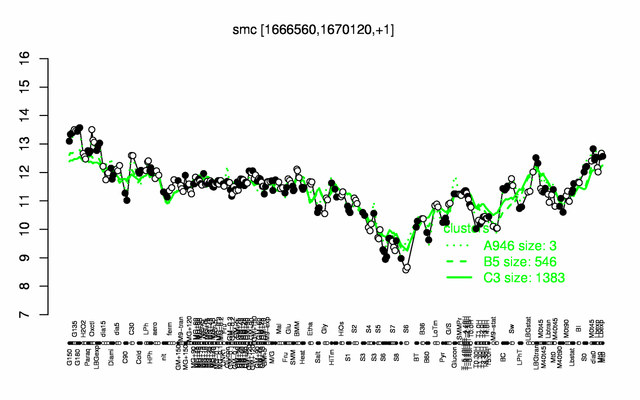
| |
Contents
Categories containing this gene/protein
DNA condensation/ segregation, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15940
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: SMC family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: nucleoid (heterogeneous) PubMed
Database entries
- Structure:
- UniProt: P51834
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
M E Fuentes-Perez, E J Gwynn, M S Dillingham, F Moreno-Herrero
Using DNA as a fiducial marker to study SMC complex interactions with the atomic force microscope.
Biophys J: 2012, 102(4);839-48
[PubMed:22385855]
[WorldCat.org]
[DOI]
(I p)
Elodie Marchadier, Rut Carballido-López, Sophie Brinster, Céline Fabret, Peggy Mervelet, Philippe Bessières, Marie-Françoise Noirot-Gros, Vincent Fromion, Philippe Noirot
An expanded protein-protein interaction network in Bacillus subtilis reveals a group of hubs: Exploration by an integrative approach.
Proteomics: 2011, 11(15);2981-91
[PubMed:21630458]
[WorldCat.org]
[DOI]
(I p)
Nora L Sullivan, Kathleen A Marquis, David Z Rudner
Recruitment of SMC by ParB-parS organizes the origin region and promotes efficient chromosome segregation.
Cell: 2009, 137(4);697-707
[PubMed:19450517]
[WorldCat.org]
[DOI]
(I p)
Stephan Gruber, Jeff Errington
Recruitment of condensin to replication origin regions by ParB/SpoOJ promotes chromosome segregation in B. subtilis.
Cell: 2009, 137(4);685-96
[PubMed:19450516]
[WorldCat.org]
[DOI]
(I p)
Robert A Britton, Elke Küster-Schöck, Thomas A Auchtung, Alan D Grossman
SOS induction in a subpopulation of structural maintenance of chromosome (Smc) mutant cells in Bacillus subtilis.
J Bacteriol: 2007, 189(12);4359-66
[PubMed:17416649]
[WorldCat.org]
[DOI]
(P p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Etienne Dervyn, Marie-Françoise Noirot-Gros, Peggy Mervelet, Steven McGovern, S Dusko Ehrlich, Patrice Polard, Philippe Noirot
The bacterial condensin/cohesin-like protein complex acts in DNA repair and regulation of gene expression.
Mol Microbiol: 2004, 51(6);1629-40
[PubMed:15009890]
[WorldCat.org]
[DOI]
(P p)
Janet C Lindow, Masayoshi Kuwano, Shigeki Moriya, Alan D Grossman
Subcellular localization of the Bacillus subtilis structural maintenance of chromosomes (SMC) protein.
Mol Microbiol: 2002, 46(4);997-1009
[PubMed:12421306]
[WorldCat.org]
[DOI]
(P p)
Jörg Soppa, Kazuo Kobayashi, Marie-Françoise Noirot-Gros, Dieter Oesterhelt, S Dusko Ehrlich, Etienne Dervyn, Naotake Ogasawara, Shigeki Moriya
Discovery of two novel families of proteins that are proposed to interact with prokaryotic SMC proteins, and characterization of the Bacillus subtilis family members ScpA and ScpB.
Mol Microbiol: 2002, 45(1);59-71
[PubMed:12100548]
[WorldCat.org]
[DOI]
(P p)
P L Graumann, R Losick
Coupling of asymmetric division to polar placement of replication origin regions in Bacillus subtilis.
J Bacteriol: 2001, 183(13);4052-60
[PubMed:11395470]
[WorldCat.org]
[DOI]
(P p)
P L Graumann, R Losick, A V Strunnikov
Subcellular localization of Bacillus subtilis SMC, a protein involved in chromosome condensation and segregation.
J Bacteriol: 1998, 180(21);5749-55
[PubMed:9791128]
[WorldCat.org]
[DOI]
(P p)