Difference between revisions of "Zwf"
(→Biological materials) |
Raphael2215 (talk | contribs) (→Database entries) |
||
| Line 81: | Line 81: | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' [http://www.pdb.org/pdb/explore/explore.do?structureId=1DPG 1DPG] (from ''Leuconostoc Mesenteroides'', 45% identity, 62% similarity) {{PubMed|7881907}} |
* '''UniProt:''' [http://www.uniprot.org/uniprot/P54547 P54547] | * '''UniProt:''' [http://www.uniprot.org/uniprot/P54547 P54547] | ||
Revision as of 10:55, 21 July 2010
- Description: glucose 6-phosphate dehydrogenase, pentose-phosphate pathway
| Gene name | zwf |
| Synonyms | yqjJ |
| Essential | no |
| Product | glucose 6-phosphate dehydrogenase |
| Function | initiation of the pentose phosphate pathway |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 55 kDa, 5.28 |
| Gene length, protein length | 1467 bp, 489 aa |
| Immediate neighbours | rnz, gndA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 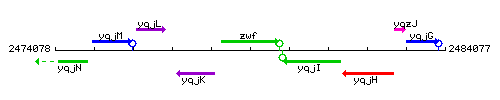
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU23850
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: D-glucose 6-phosphate + NADP+ = D-glucono-1,5-lactone 6-phosphate + NADPH (according to Swiss-Prot)
- Protein family: glucose-6-phosphate dehydrogenase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: Two Substrate Michaelis-Menten PubMed
- Domains:
- Modification:
- Cofactor(s): Mg2+, Mn2+, Ca2+
- Effectors of protein activity:
- NAD+, NADP+ and NADPH affect the enzyme kinetic PubMed
- Interactions:
- Localization:
Database entries
- UniProt: P54547
- KEGG entry: [3]
- E.C. number: 1.1.1.49
Additional information
The enzyme is a dimer PubMed
Expression and regulation
- Operon: zwf PubMed
- Regulation: constitutive PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- SM-NB3 (zwf-spc), available in Anne Galinier's and Boris Görke's labs
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Yun Xia Duan, Tao Chen, Xun Chen, Xue Ming Zhao
Overexpression of glucose-6-phosphate dehydrogenase enhances riboflavin production in Bacillus subtilis.
Appl Microbiol Biotechnol: 2010, 85(6);1907-14
[PubMed:19779711]
[WorldCat.org]
[DOI]
(I p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
S Ujita, K Kimura
Glucose-6-phosphate dehydrogenase, vegetative and spore Bacillus subtilis.
Methods Enzymol: 1982, 89 Pt D;258-61
[PubMed:6292660]
[WorldCat.org]
[DOI]
(P p)