Difference between revisions of "SigL"
| Line 124: | Line 124: | ||
=References= | =References= | ||
| − | + | ||
# Ali, N. O., Bignon, J., Rapoport, G., and Débarbouillé, M. (2001) Regulation of the acetoin catabolic pathway is controlled by sigma L in Bacillus subtilis. J. Bacteriol. 183, 2497-2504. [http://www.ncbi.nlm.nih.gov/sites/entrez/11274109 PubMed] | # Ali, N. O., Bignon, J., Rapoport, G., and Débarbouillé, M. (2001) Regulation of the acetoin catabolic pathway is controlled by sigma L in Bacillus subtilis. J. Bacteriol. 183, 2497-2504. [http://www.ncbi.nlm.nih.gov/sites/entrez/11274109 PubMed] | ||
# Belitsky BR, Sonenshein AL (1998) Role and regulation of Bacillus subtilis glutamate dehydrogenase genes. J Bacteriol 180:6298-6305. [http://www.ncbi.nlm.nih.gov/sites/entrez/9829940 PubMed] | # Belitsky BR, Sonenshein AL (1998) Role and regulation of Bacillus subtilis glutamate dehydrogenase genes. J Bacteriol 180:6298-6305. [http://www.ncbi.nlm.nih.gov/sites/entrez/9829940 PubMed] | ||
Revision as of 00:52, 3 April 2009
- Description: Sigma subunit of the RNA polymerase, Sigma-54, Sigma L
| Gene name | sigL |
| Synonyms | |
| Essential | no |
| Product | RNA polymerase sigma-54 factor (sigma-L) |
| Function | utilization of arginin, acetoin, and fructose, required for cold adaptation |
| MW, pI | 49,5 kDa, 7.79 |
| Gene length, protein length | 1308 bp, 436 amino acids |
| Immediate neighbours | yvfG, yvfH |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 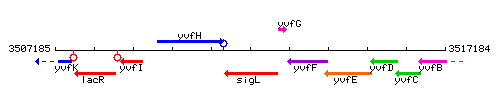
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates: 3511532 - 3512839
Phenotypes of a mutant
The mutant is cold-sensitive and unable to use arginine as a single carbon source. PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Binding to promoters of the -12, -24 type
- Protein family: sigma-54 factor family
- Paralogous protein(s):
Genes/ operons controlled by SigL
acoA-acoB-acoC-acoL, rocA-rocB-rocC, rocD-rocE-rocF, rocG, levD-levE-levF-levG-sacC, ptb-bcd-buk-lpdV-bkdAA-bkdAB-bkdB
Extended information on the protein
- Kinetic information:
- Domains:
- DNA binding domain (H-T-H motif) (324–343)
- pron box domain (413–421)
- 3 x Compositional bias domain (6–21),(32–53),(112–136)
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- Structure:
- Swiss prot entry: [3]
- KEGG entry: [4]
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulatory mechanism: CcpA: transcription repression, transcriptional roadblock
- Additional information:
Biological materials
- Mutant: GP146 (aphA3), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- Antibody:
Labs working on this gene/protein
Michel Debarbouille, Pasteur Institute, Paris, France Homepage
Your additional remarks
References
- Ali, N. O., Bignon, J., Rapoport, G., and Débarbouillé, M. (2001) Regulation of the acetoin catabolic pathway is controlled by sigma L in Bacillus subtilis. J. Bacteriol. 183, 2497-2504. PubMed
- Belitsky BR, Sonenshein AL (1998) Role and regulation of Bacillus subtilis glutamate dehydrogenase genes. J Bacteriol 180:6298-6305. PubMed
- Choi SK, Saier MH Jr: (2005) Regulation of sigL expression by the catabolite control protein CcpA involves a roadblock mechanism in Bacillus subtilis: potential connection between carbon and nitrogen metabolism. J Bacteriol , 187:6856-6861. PubMed
- Débarbouillé M, Martin-Verstraete I, Kunst F, Rapoport G: (1991) The Bacillus subtilis sigL gene encodes an equivalent of (Sigma)54 from Gram-negative bacteria. Proc Natl Acad Sci USA , 88:9092-9096. PubMed
- Martin-Verstraete I, Débarbouillé M, Klier A, Rapoport G: (1992) Mutagenesis of the Bacillus subtilis ‘-12, -24’ promoter of the levanase operon and evidence for the existence of an upstream activating sequence. J Mol Biol , 226:85-99. PubMed
- Wiegeshoff et al. (2006) Sigma L is important for cold shock adaptation of Bacillus subtilis. J Bacteriol , 188: 3130-3133. PubMed