Difference between revisions of "Rny"
| Line 86: | Line 86: | ||
* '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/O31774] | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/O31774] | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU16960] |
* '''E.C. number:''' [http://www.expasy.org/enzyme/3.1.4.16] | * '''E.C. number:''' [http://www.expasy.org/enzyme/3.1.4.16] | ||
Revision as of 12:50, 20 March 2009
- Description: RNase Y, endoribonuclease, required for the processing of the gapA operon mRNA
| Gene name | rny |
| Synonyms | ymdA |
| Essential | yes |
| Product | RNase Y |
| Function | required for the processing of the gapA operon mRNA |
| MW, pI | 58,7 kDa, 5.39 |
| Gene length, protein length | 1560 bp, 520 amino acids |
| Immediate neighbours | pbpX, ymdB |
| Gene sequence (+200bp) corrected | Protein sequence |
| Caution: The sequence for this gene in SubtiList contains errors | |
Genetic context 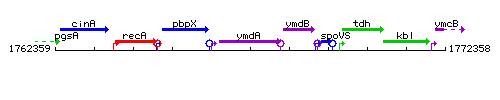
| |
Contents
The gene
Basic information
- Coordinates: 1766580 - 1768139
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Protein family: 2',3' cyclic nucleotide phosphodiesterase family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- transmembrane domain (4–24)
- KH domain (210–273)
- HD domain (336–429)
- Modification:
- Cofactor(s): metal-dependent
- Effectors of protein activity:
- Localization: Single-pass membrane protein PubMed
Database entries
- Structure:
- Swiss prot entry: [2]
- KEGG entry: [3]
- E.C. number: [4]
Additional information
required for the processing of the gapA operon mRNA
Expression and regulation
- Sigma factor:
- Regulation: constitutive
- Regulatory mechanism:
- Additional information: there is a terminator between rny and ymdB, most transcripts terminate there
Biological materials
- Mutant: essential!!!!, 4043 (rny under p-spac control), GP193 (rny under p-xyl control), available in Stülke lab
- Expression vector:
N-terminal Strep-tag, expression in E. coli, in pGP172: pGP441, available in Stülke lab
N-terminal Strep-tag, for SPINE, expression in B. subtilis, in pGP380: pGP775 , available in Stülke lab
wild type rny, expression in B. subtilis, in pBQ200: pGP1201, available in Stülke lab
there is also a series of domain constructs present in pBQ200, all available in Stülke lab
- GFP fusion: B. subtilis 3569 (amyE:: (p-xyl rny-gfpmut1-spc)), available in Errington lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
Labs working on this gene/protein
Harald Putzer, IBPC Paris, France Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
- Commichau, F. M., Rothe, F. M., Herzberg, C., Wagner, E., Hellwig, D., Lehnik-Habrink, M., Hammer, E., Völker, U. & Stülke, J. (2009) Novel activities of glycolytic enzymes in Bacillus subtilis: Interactions with essential proteins involved in mRNA processing. Mol. Cell. Proteomics in press PubMed
- Hunt, A., Rawlins, J. P., Thomaides, H. B., and Errington, J. (2006) Functional analysis of 11 putative essential genes in Bacillus subtilis. Microbiology 152, 2895-2907. PubMed