Difference between revisions of "ScpB"
(→Original publications) |
(→Original publications) |
||
| Line 148: | Line 148: | ||
<pubmed> 22933559 22934648 24118085 </pubmed> | <pubmed> 22933559 22934648 24118085 </pubmed> | ||
== Original publications == | == Original publications == | ||
| − | <pubmed>12100548,12065423,12421306,22385855, 16479537, 19450516 12897137 7934830 11948165 23353789 24440399,24440393 25071173, 25557547 25951515 26253537 23475963</pubmed> | + | <pubmed>12100548,12065423,12421306,22385855, 16479537, 19450516 12897137 7934830 11948165 23353789 24440399,24440393 25071173, 25557547 25951515 26253537 23475963 26295962</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 08:53, 24 August 2015
- Description: part of the condensin complex, chromosomal origin condensation and segregation
| Gene name | scpB |
| Synonyms | ypuH |
| Essential | yes PubMed |
| Product | DNA segregation and condensation protein |
| Function | segregation of replication origins |
| Gene expression levels in SubtiExpress: scpB | |
| Interactions involving this protein in SubtInteract: ScpB | |
| MW, pI | 21 kDa, 4.25 |
| Gene length, protein length | 591 bp, 197 aa |
| Immediate neighbours | ypuI, scpA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 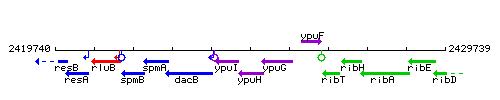
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed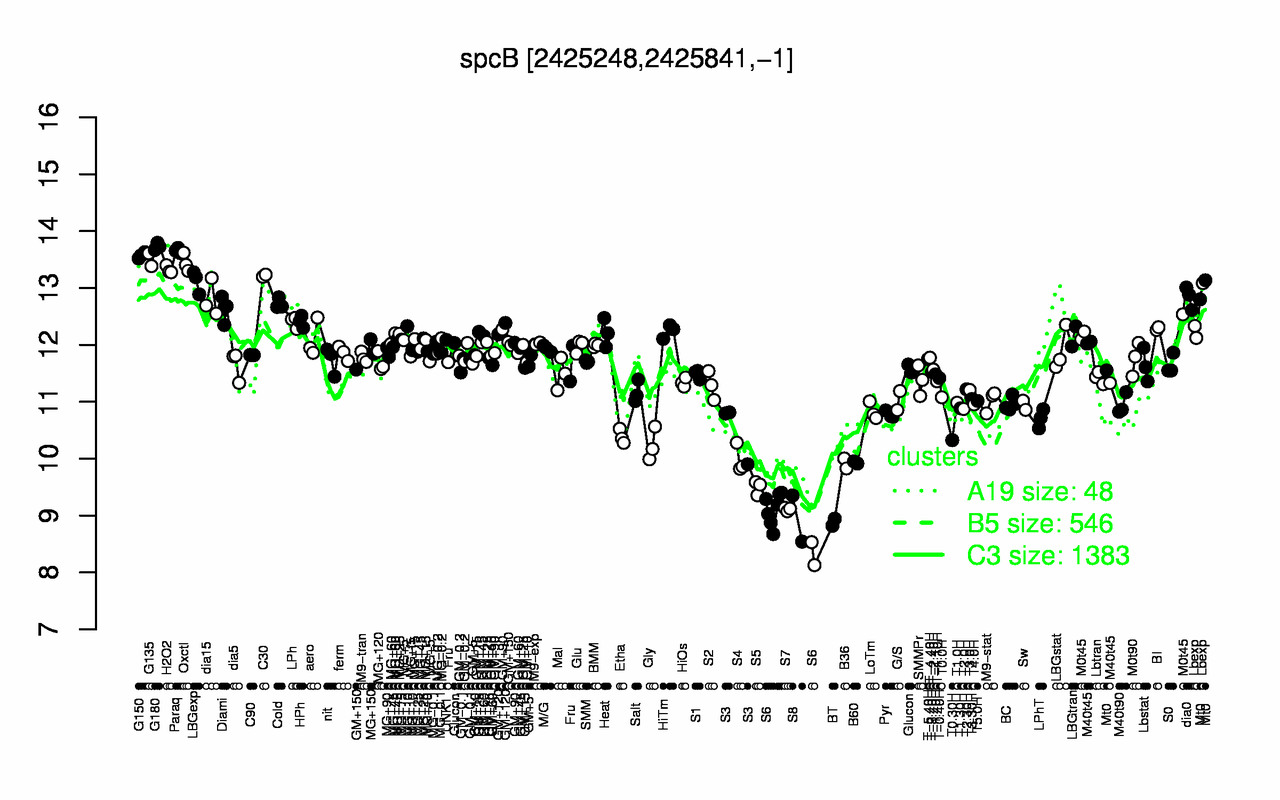
| |
Contents
Categories containing this gene/protein
DNA condensation/ segregation, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23210
Phenotypes of a mutant
- essential PubMed
- scpB mutants are not viable on complex medim that allow rapid growth, but they are viable under conditions of slow growth PubMed
Database entries
- BsubCyc: BSU23210
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: scpB family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- BsubCyc: BSU23210
- UniProt: P35155
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications