Difference between revisions of "YjbC"
(→References) |
|||
| Line 1: | Line 1: | ||
| − | * '''Description:''' general stress protein, required for survival of salt | + | * '''Description:''' general stress protein, required for survival of salt and paraquat stresses <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
| Line 11: | Line 11: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || unknown | |style="background:#ABCDEF;" align="center"| '''Product''' || unknown | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || survival of paraquat stress |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU11490 yjbC] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU11490 yjbC] | ||
| Line 36: | Line 36: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/> | <br/><br/> | ||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
{{SubtiWiki category|[[general stress proteins (controlled by SigB)]]}}, | {{SubtiWiki category|[[general stress proteins (controlled by SigB)]]}}, | ||
| + | {{SubtiWiki category|[[resistance against oxidative and electrophile stress]]}}, | ||
{{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}} | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}} | ||
| Line 118: | Line 115: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yjbC_1226938_1227516_1 yjbC] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yjbC_1226938_1227516_1 yjbC] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' [[SigW]], [[SigB]], [[SigX]], [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+10913081 PubMed], [[SigM]] [http://www.ncbi.nlm.nih.gov/sites/entrez/17434969 PubMed] | + | * '''[[Sigma factor]]:''' [[SigW]], [[SigB]], [[SigX]], [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+10913081 PubMed], [[SigM]] [http://www.ncbi.nlm.nih.gov/sites/entrez/17434969 PubMed] |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 150: | Line 147: | ||
=References= | =References= | ||
| − | <pubmed>10913081,,17158660,15805528 17158660 15805528,21815947 </pubmed> | + | <pubmed>10913081,22582280,17158660,15805528 17158660 15805528,21815947 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 18:54, 15 September 2013
- Description: general stress protein, required for survival of salt and paraquat stresses
| Gene name | yjbC |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | survival of paraquat stress |
| Gene expression levels in SubtiExpress: yjbC | |
| MW, pI | 22 kDa, 5.14 |
| Gene length, protein length | 576 bp, 192 aa |
| Immediate neighbours | yjbB, spx |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 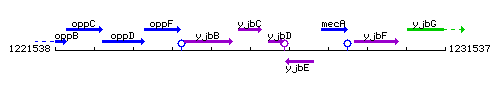
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed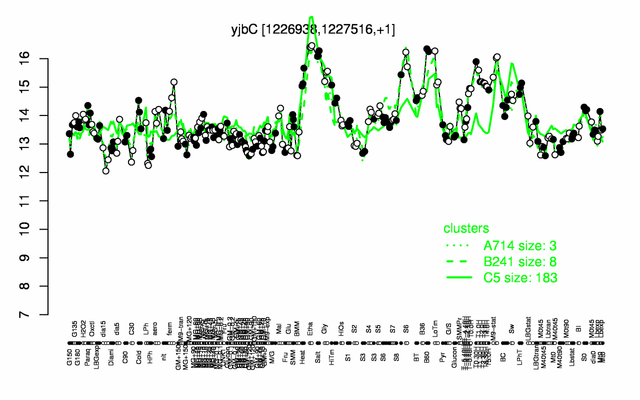
| |
Contents
Categories containing this gene/protein
general stress proteins (controlled by SigB), resistance against oxidative and electrophile stress, cell envelope stress proteins (controlled by SigM, V, W, X, Y)
This gene is a member of the following regulons
PerR regulon, SigB regulon, SigM regulon, SigW regulon, SigX regulon
The gene
Basic information
- Locus tag: BSU11490
Phenotypes of a mutant
strongly impaired survival of salt stress PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O31601
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References