Difference between revisions of "LicT"
| Line 33: | Line 33: | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=licT_4012866_4013699_-1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:licT_expression.png|500px]] | + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=licT_4012866_4013699_-1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:licT_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU39080]] |
|- | |- | ||
|} | |} | ||
Revision as of 14:57, 16 May 2013
- Description: transcriptional antiterminator of the bglP-bglH-yxiE operon and the bglS gene
| Gene name | licT |
| Synonyms | |
| Essential | no |
| Product | transcriptional antiterminator (BglG family) |
| Function | control of beta-glucan and beta-glucoside utilization |
| Gene expression levels in SubtiExpress: licT | |
| Interactions involving this protein in SubtInteract: LicT | |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 32 kDa, 5.944 |
| Gene length, protein length | 831 bp, 277 aa |
| Immediate neighbours | bglS, yxiP |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 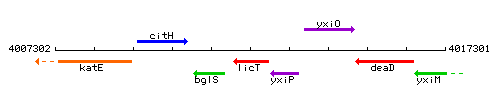
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed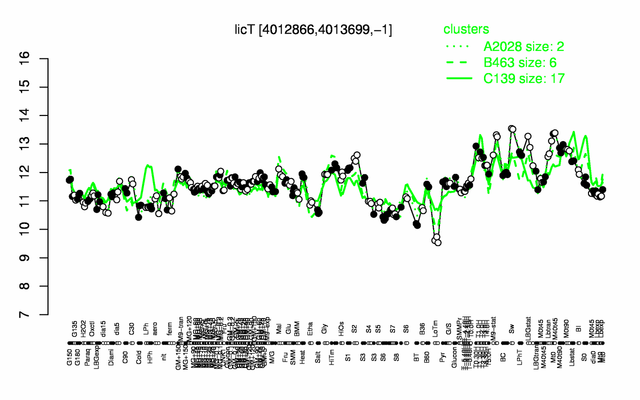
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, transcription factors and their control, RNA binding regulators, phosphoproteins
This gene is a member of the following regulons
The LicT regulon: bglP-bglH-yxiE, bglS
The gene
Basic information
- Locus tag: BSU39080
Phenotypes of a mutant
no expression of the bglP-bglH operon
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: binding to the mRNAs of bglS and the bglP-bglH operon, causes transcription antitermination (in presence of salicin and absence of glucose)
- Protein family: transcriptional antiterminator BglG family of antiterminators (according to Swiss-Prot)
Extended information on the protein
- Kinetic information:
- K(D) for the RAT-RNA: 10 nM PubMed
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm, even distribution in the absence of the inducer salicin, subpolar localization in the presence of salicin PubMed
Database entries
- UniProt: P39805
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP427 (licTS, erm), available in the Stülke lab
- Expression vector:
- for expression, purification of both PRDs in E. coli with N-terminal His-tag, in pWH844: pGP165, available in Stülke lab
- for expression, purification of the RNA-binding domain in E. coli with N-terminal His-tag, in pWH844: pGP315, available in Stülke lab
- for expression, purification of the RNA-binding domain in E. coli with N-terminal His-tag and thrombin cleavage site, in pGP570: pGP572, available in Stülke lab
- lacZ fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Josef Deutscher, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Michael Hecker, Greifswald, Germany Homepage
Your additional remarks
References
Original description
K Schnetz, J Stülke, S Gertz, S Krüger, M Krieg, M Hecker, B Rak
LicT, a Bacillus subtilis transcriptional antiterminator protein of the BglG family.
J Bacteriol: 1996, 178(7);1971-9
[PubMed:8606172]
[WorldCat.org]
[DOI]
(P p)
Control of LicT activity
Structural analysis of LicT
Hélène Déméné, Thierry Ducat, Karine De Guillen, Catherine Birck, Stéphane Aymerich, Michel Kochoyan, Nathalie Declerck
Structural mechanism of signal transduction between the RNA-binding domain and the phosphotransferase system regulation domain of the LicT antiterminator.
J Biol Chem: 2008, 283(45);30838-49
[PubMed:18682383]
[WorldCat.org]
[DOI]
(P p)
Marc Graille, Cong-Zhao Zhou, Véronique Receveur-Bréchot, Bruno Collinet, Nathalie Declerck, Herman van Tilbeurgh
Activation of the LicT transcriptional antiterminator involves a domain swing/lock mechanism provoking massive structural changes.
J Biol Chem: 2005, 280(15);14780-9
[PubMed:15699035]
[WorldCat.org]
[DOI]
(P p)
N Declerck, H Dutartre, V Receveur, V Dubois, C Royer, S Aymerich, H van Tilbeurgh
Dimer stabilization upon activation of the transcriptional antiterminator LicT.
J Mol Biol: 2001, 314(4);671-81
[PubMed:11733988]
[WorldCat.org]
[DOI]
(P p)
H van Tilbeurgh, D Le Coq, N Declerck
Crystal structure of an activated form of the PTS regulation domain from the LicT transcriptional antiterminator.
EMBO J: 2001, 20(14);3789-99
[PubMed:11447120]
[WorldCat.org]
[DOI]
(P p)
LicT-RNA interaction
Caroline Clerte, Nathalie Declerck, Emmanuel Margeat
Competitive folding of anti-terminator/terminator hairpins monitored by single molecule FRET.
Nucleic Acids Res: 2013, 41(4);2632-43
[PubMed:23303779]
[WorldCat.org]
[DOI]
(I p)
Sebastian Hübner, Nathalie Declerck, Christine Diethmaier, Dominique Le Coq, Stephane Aymerich, Jörg Stülke
Prevention of cross-talk in conserved regulatory systems: identification of specificity determinants in RNA-binding anti-termination proteins of the BglG family.
Nucleic Acids Res: 2011, 39(10);4360-72
[PubMed:21278164]
[WorldCat.org]
[DOI]
(I p)
Hélène Déméné, Thierry Ducat, Karine De Guillen, Catherine Birck, Stéphane Aymerich, Michel Kochoyan, Nathalie Declerck
Structural mechanism of signal transduction between the RNA-binding domain and the phosphotransferase system regulation domain of the LicT antiterminator.
J Biol Chem: 2008, 283(45);30838-49
[PubMed:18682383]
[WorldCat.org]
[DOI]
(P p)
Yinshan Yang, Nathalie Declerck, Xavier Manival, Stéphane Aymerich, Michel Kochoyan
Solution structure of the LicT-RNA antitermination complex: CAT clamping RAT.
EMBO J: 2002, 21(8);1987-97
[PubMed:11953318]
[WorldCat.org]
[DOI]
(P p)
N Declerck, F Vincent, F Hoh, S Aymerich, H van Tilbeurgh
RNA recognition by transcriptional antiterminators of the BglG/SacY family: functional and structural comparison of the CAT domain from SacY and LicT.
J Mol Biol: 1999, 294(2);389-402
[PubMed:10610766]
[WorldCat.org]
[DOI]
(P p)
S Aymerich, M Steinmetz
Specificity determinants and structural features in the RNA target of the bacterial antiterminator proteins of the BglG/SacY family.
Proc Natl Acad Sci U S A: 1992, 89(21);10410-4
[PubMed:1279678]
[WorldCat.org]
[DOI]
(P p)