Difference between revisions of "SsuD"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 44: | Line 44: | ||
* '''Locus tag:''' BSU08860 | * '''Locus tag:''' BSU08860 | ||
| − | |||
| − | |||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
Revision as of 20:34, 27 January 2012
- Description: aliphatic sulfonate monooxygenase
| Gene name | ssuD |
| Synonyms | ygcA, yzeC |
| Essential | no |
| Product | aliphatic sulfonate monooxygenase |
| Function | sulfonate reduction |
| Metabolic function and regulation of this protein in SubtiPathways: Cys, Met & Sulfate assimilation | |
| MW, pI | 41 kDa, 5.369 |
| Gene length, protein length | 1128 bp, 376 aa |
| Immediate neighbours | ssuC, ygaN |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 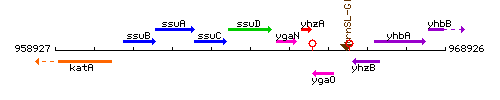
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU08860
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: An alkanesufonate (R-CH(2)-SO3H) + FMNH2 + O2 = an aldehyde (R-CHO) + FMN + sulfite + H2O (according to Swiss-Prot)
- Protein family: ssuD family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 1NQK (from Escherichia coli, 64% identity, 76% similarity)
- UniProt: P40402
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Sergine Even, Pierre Burguière, Sandrine Auger, Olga Soutourina, Antoine Danchin, Isabelle Martin-Verstraete
Global control of cysteine metabolism by CymR in Bacillus subtilis.
J Bacteriol: 2006, 188(6);2184-97
[PubMed:16513748]
[WorldCat.org]
[DOI]
(P p)
Kyle N Erwin, Shunji Nakano, Peter Zuber
Sulfate-dependent repression of genes that function in organosulfur metabolism in Bacillus subtilis requires Spx.
J Bacteriol: 2005, 187(12);4042-9
[PubMed:15937167]
[WorldCat.org]
[DOI]
(P p)
J R van der Ploeg, M Barone, T Leisinger
Expression of the Bacillus subtilis sulphonate-sulphur utilization genes is regulated at the levels of transcription initiation and termination.
Mol Microbiol: 2001, 39(5);1356-65
[PubMed:11251850]
[WorldCat.org]
(P p)
Jan R van der Ploeg, Nicola J Cummings, Thomas Leisinger, Ian F Connerton
Bacillus subtilis genes for the utilization of sulfur from aliphatic sulfonates.
Microbiology (Reading): 1998, 144 ( Pt 9);2555-2561
[PubMed:9782504]
[WorldCat.org]
[DOI]
(P p)