Difference between revisions of "AlaT"
(→Extended information on the protein) |
|||
| Line 75: | Line 75: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
* '''Localization:''' | * '''Localization:''' | ||
Revision as of 16:57, 13 July 2011
- Description: similar to PLP-dependent methionine aminotransferase
| Gene name | alaT |
| Synonyms | yugH |
| Essential | no |
| Product | unknown |
| Function | unknown |
| MW, pI | 42 kDa, 5.106 |
| Gene length, protein length | 1158 bp, 386 aa |
| Immediate neighbours | yugI, alaR |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 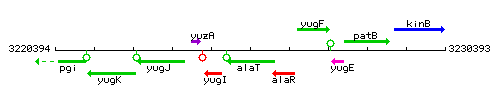
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
poorly characterized/ putative enzymes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU31400
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- Structure:
- UniProt: Q795M6
- KEGG entry: [2]
- E.C. number:
Additional information
The gene is annotated in KEGG as an ortholog of N-succinyldiaminopimelate aminotransferase EC 2.6.1.17. It is annotated in Swiss-Prot as “alanine transaminase”. In MetaCyc the gene is marked as “similar to aspartate aminotransferase”. No literature/experimental evidence supporting the annotation is available. PubMed
Expression and regulation
- Regulation:
- Regulatory mechanism:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Irnov Irnov, Cynthia M Sharma, Jörg Vogel, Wade C Winkler
Identification of regulatory RNAs in Bacillus subtilis.
Nucleic Acids Res: 2010, 38(19);6637-51
[PubMed:20525796]
[WorldCat.org]
[DOI]
(I p)