Difference between revisions of "CypC"
(→References) |
(→References) |
||
| Line 128: | Line 128: | ||
=References= | =References= | ||
'''Additional publications:''' {{PubMed|17385817,11914497}} | '''Additional publications:''' {{PubMed|17385817,11914497}} | ||
| − | <pubmed>,10529095,12519760,15805528, 20697922 | + | <pubmed>12297285 11827534 11566026 17385817,11914497</pubmed> |
| + | --- | ||
| + | <pubmed>,10529095,12519760,15805528, 20697922 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 15:21, 16 April 2011
- Description: long chain-fatty acid beta-hydroxylating cytochrome P450, H(2)O(2)-dependent, hydroxylates myristic acid to beta-hydroxymyristic acid
| Gene name | cypC |
| Synonyms | ybdT |
| Essential | no |
| Product | fatty acid beta-hydroxylating cytochrome P450 |
| Function | biosynthesis of beta-hydroxy fatty acid |
| MW, pI | 47 kDa, 6.468 |
| Gene length, protein length | 1251 bp, 417 aa |
| Immediate neighbours | ybxI, ybyB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 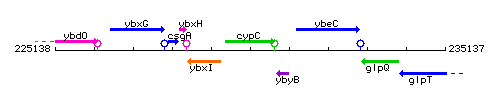
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
electron transport/ other, lipid metabolism/ other
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU02100
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- hydroxylation of a long-chain fatty acid (e.g. myristic acid) at the alpha- and beta-positions using hydrogen peroxide as an oxidant PubMed
- Protein family: cytochrome P450 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- UniProt: O31440
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: cypC (according to DBTBS)
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
---