Difference between revisions of "YurJ"
| Line 30: | Line 30: | ||
<br/><br/> | <br/><br/> | ||
| + | |||
| + | = [[Categories]] containing this gene/protein = | ||
| + | {{SubtiWiki category|[[ABC transporters]]}}, | ||
| + | {{SubtiWiki category|[[utilization of specific carbon sources]]}}, | ||
| + | {{SubtiWiki category|[[utilization of nitrogen sources other than amino acids]]}}, | ||
| + | {{SubtiWiki category|[[membrane proteins]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| + | {{SubtiWiki regulon|[[CodY regulon]]}} | ||
=The gene= | =The gene= | ||
| Line 49: | Line 58: | ||
| − | + | ||
| − | |||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
Revision as of 23:08, 8 December 2010
- Description: aminosugar ABC transporter (ATP-binding protein)
| Gene name | yurJ |
| Synonyms | |
| Essential | no |
| Product | aminosugar ABC transporter (ATP-binding protein) |
| Function | uptake of sugar amines |
| MW, pI | 41 kDa, 6.071 |
| Gene length, protein length | 1101 bp, 367 aa |
| Immediate neighbours | yurI, yurK |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 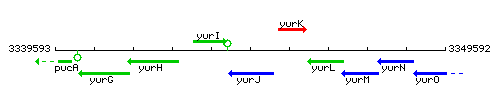
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
ABC transporters, utilization of specific carbon sources, utilization of nitrogen sources other than amino acids, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU32550
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylated on Ser-3 PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O32151
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Boumediene Soufi, Chanchal Kumar, Florian Gnad, Matthias Mann, Ivan Mijakovic, Boris Macek
Stable isotope labeling by amino acids in cell culture (SILAC) applied to quantitative proteomics of Bacillus subtilis.
J Proteome Res: 2010, 9(7);3638-46
[PubMed:20509597]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Virginie Molle, Yoshiko Nakaura, Robert P Shivers, Hirotake Yamaguchi, Richard Losick, Yasutaro Fujita, Abraham L Sonenshein
Additional targets of the Bacillus subtilis global regulator CodY identified by chromatin immunoprecipitation and genome-wide transcript analysis.
J Bacteriol: 2003, 185(6);1911-22
[PubMed:12618455]
[WorldCat.org]
[DOI]
(P p)
Y Quentin, G Fichant, F Denizot
Inventory, assembly and analysis of Bacillus subtilis ABC transport systems.
J Mol Biol: 1999, 287(3);467-84
[PubMed:10092453]
[WorldCat.org]
[DOI]
(P p)