Difference between revisions of "MurAA"
(→Basic information/ Evolution) |
|||
| Line 52: | Line 52: | ||
| + | |||
| + | = Categories containing this gene/protein = | ||
| + | {{SubtiWiki category|[[cell wall synthesis]]}}, | ||
| + | {{SubtiWiki category|[[biosynthesis of cell wall components]]}}, | ||
| + | {{SubtiWiki category|[[essential genes]]}} | ||
=The protein= | =The protein= | ||
Revision as of 18:15, 30 November 2010
- Description: UDP-N-acetylglucosamine 1-carboxyvinyltransferase
| Gene name | murAA |
| Synonyms | murA |
| Essential | yes PubMed |
| Product | UDP-N-acetylglucosamine 1-carboxyvinyltransferase |
| Function | peptidoglycan precursor biosynthesis |
| Metabolic function and regulation of this protein in SubtiPathways: Cell wall | |
| MW, pI | 46 kDa, 5.45 |
| Gene length, protein length | 1308 bp, 436 aa |
| Immediate neighbours | spoIID, ywmB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 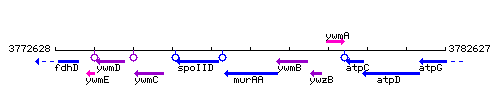
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU36760
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
Categories containing this gene/protein
cell wall synthesis, biosynthesis of cell wall components, essential genes
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Phosphoenolpyruvate + UDP-N-acetyl-D-glucosamine = phosphate + UDP-N-acetyl-3-O-(1-carboxyvinyl)-D-glucosamine (according to Swiss-Prot)
- Protein family: MurA subfamily (according to Swiss-Prot)
- Paralogous protein(s): MurAB
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- UniProt: P70965
- KEGG entry: [3]
- E.C. number: 2.5.1.7
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References