Difference between revisions of "NagZ"
(→References) |
|||
| Line 38: | Line 38: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| + | increased autolysis {{PubMed|20400549}} | ||
=== Database entries === | === Database entries === | ||
| Line 52: | Line 53: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' cleaves muropeptides derived from peptidoglycan, but not peptidoglycan itself {{PubMed|20400549}} |
* '''Protein family:''' glycosyl hydrolase 3 family (according to Swiss-Prot) | * '''Protein family:''' glycosyl hydrolase 3 family (according to Swiss-Prot) | ||
| Line 72: | Line 73: | ||
* '''Interactions:''' | * '''Interactions:''' | ||
| − | * '''Localization:''' | + | * '''Localization:''' secreted (with signal peptide), remains to some extent cell wall-associated {{PubMed|20400549}} |
=== Database entries === | === Database entries === | ||
| Line 85: | Line 86: | ||
=== Additional information=== | === Additional information=== | ||
| − | The gene is annotated in KEGG as an ortholog of beta-N-acetylhexosaminidase EC 3.2.1.52. It is marked in MetaCyc as “similar to beta-hexosaminidase”. No EC annotation is available in Swiss-ProtSwiss-Prot.supporting the annotation is available. {{PubMed|19935659}} | + | The gene is mis-annotated in KEGG as an ortholog of beta-N-acetylhexosaminidase EC 3.2.1.52. It is marked in MetaCyc as “similar to beta-hexosaminidase”. No EC annotation is available in Swiss-ProtSwiss-Prot.supporting the annotation is available. {{PubMed|19935659}} |
=Expression and regulation= | =Expression and regulation= | ||
| Line 94: | Line 95: | ||
* '''Regulation:''' | * '''Regulation:''' | ||
| + | ** expressed in late exponential and early stationary phase {{PubMed|20400549}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| Line 101: | Line 103: | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' available in [[Christoph Mayer]]'s lab {{PubMed|20400549}} |
* '''Expression vector:''' | * '''Expression vector:''' | ||
Revision as of 16:50, 20 April 2010
- Description: N-acetylglucosaminidase
| Gene name | ybbD |
| Synonyms | yzbA |
| Essential | no |
| Product | N-acetylglucosaminidase |
| Function | cell wall recycling |
| MW, pI | 70 kDa, 9.76 |
| Gene length, protein length | 1926 bp, 642 aa |
| Immediate neighbours | ybbC, ybbE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 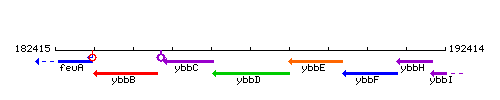
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU01660
Phenotypes of a mutant
increased autolysis PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: cleaves muropeptides derived from peptidoglycan, but not peptidoglycan itself PubMed
- Protein family: glycosyl hydrolase 3 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: secreted (with signal peptide), remains to some extent cell wall-associated PubMed
Database entries
- Structure: 3CQM
- UniProt: P40406
- KEGG entry: [2]
- E.C. number:
Additional information
The gene is mis-annotated in KEGG as an ortholog of beta-N-acetylhexosaminidase EC 3.2.1.52. It is marked in MetaCyc as “similar to beta-hexosaminidase”. No EC annotation is available in Swiss-ProtSwiss-Prot.supporting the annotation is available. PubMed
Expression and regulation
- Operon:
- Regulation:
- expressed in late exponential and early stationary phase PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: available in Christoph Mayer's lab PubMed
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References