Difference between revisions of "CpgA"
| Line 1: | Line 1: | ||
| − | * '''Description:''' GTPase, activity stimulated by ribosomes <br/><br/> | + | * '''Description:''' GTPase, activity stimulated by ribosomes, may be involved in ribosome maturation <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || GTPase | |style="background:#ABCDEF;" align="center"| '''Product''' || GTPase | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || coordination of peptidoglycan deposition in the cell wall |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/carbon_flow.html Central C-metabolism]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/carbon_flow.html Central C-metabolism]''' | ||
Revision as of 20:58, 30 October 2009
- Description: GTPase, activity stimulated by ribosomes, may be involved in ribosome maturation
| Gene name | cpgA |
| Synonyms | yloQ |
| Essential | no |
| Product | GTPase |
| Function | coordination of peptidoglycan deposition in the cell wall |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 33 kDa, 4.743 |
| Gene length, protein length | 894 bp, 298 aa |
| Immediate neighbours | prkC, rpe |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 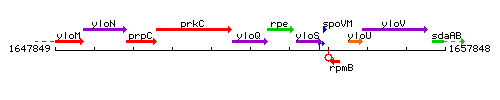
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU15780
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: engC GTPase domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure: 1T9H
- UniProt: O34530
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Tony Wilkinson, York University, U.K. homepage
Your additional remarks
References
Cédric Absalon, Michal Obuchowski, Edwige Madec, Delphine Delattre, I Barry Holland, Simone J Séror
CpgA, EF-Tu and the stressosome protein YezB are substrates of the Ser/Thr kinase/phosphatase couple, PrkC/PrpC, in Bacillus subtilis.
Microbiology (Reading): 2009, 155(Pt 3);932-943
[PubMed:19246764]
[WorldCat.org]
[DOI]
(P p)
Ishita M Shah, Maria-Halima Laaberki, David L Popham, Jonathan Dworkin
A eukaryotic-like Ser/Thr kinase signals bacteria to exit dormancy in response to peptidoglycan fragments.
Cell: 2008, 135(3);486-96
[PubMed:18984160]
[WorldCat.org]
[DOI]
(I p)
Cédric Absalon, Kassem Hamze, Didier Blanot, Claude Frehel, Rut Carballido-Lopez, Barry I Holland, Jean van Heijenoort, Simone J Séror
The GTPase CpgA is implicated in the deposition of the peptidoglycan sacculus in Bacillus subtilis.
J Bacteriol: 2008, 190(10);3786-90
[PubMed:18344364]
[WorldCat.org]
[DOI]
(I p)
Alison Hunt, Joy P Rawlins, Helena B Thomaides, Jeff Errington
Functional analysis of 11 putative essential genes in Bacillus subtilis.
Microbiology (Reading): 2006, 152(Pt 10);2895-2907
[PubMed:17005971]
[WorldCat.org]
[DOI]
(P p)
Lionel Cladière, Kassem Hamze, Edwige Madec, Vladimir M Levdikov, Anthony J Wilkinson, I Barry Holland, Simone J Séror
The GTPase, CpgA(YloQ), a putative translation factor, is implicated in morphogenesis in Bacillus subtilis.
Mol Genet Genomics: 2006, 275(4);409-20
[PubMed:16485133]
[WorldCat.org]
[DOI]
(P p)
Adam Iwanicki, Krzysztof Hinc, Simone Seror, Grzegorz Wegrzyn, Michal Obuchowski
Transcription in the prpC-yloQ region in Bacillus subtilis.
Arch Microbiol: 2005, 183(6);421-30
[PubMed:16025310]
[WorldCat.org]
[DOI]
(P p)
Tracey L Campbell, Denis M Daigle, Eric D Brown
Characterization of the Bacillus subtilis GTPase YloQ and its role in ribosome function.
Biochem J: 2005, 389(Pt 3);843-52
[PubMed:15828870]
[WorldCat.org]
[DOI]
(I p)
Vladimir M Levdikov, Elena V Blagova, James A Brannigan, Lionel Cladière, Alfred A Antson, Michail N Isupov, Simone J Séror, Anthony J Wilkinson
The crystal structure of YloQ, a circularly permuted GTPase essential for Bacillus subtilis viability.
J Mol Biol: 2004, 340(4);767-82
[PubMed:15223319]
[WorldCat.org]
[DOI]
(P p)
Lionel Cladière, Elena Blagova, Vladimir M Levdikov, James A Brannigan, Simone J Séror, Anthony J Wilkinson
Crystallization of YloQ, a GTPase of unknown function essential for Bacillus subtilis viability.
Acta Crystallogr D Biol Crystallogr: 2004, 60(Pt 2);329-30
[PubMed:14747714]
[WorldCat.org]
[DOI]
(P p)