Difference between revisions of "SacX"
| Line 1: | Line 1: | ||
| − | * '''Description:''' trigger enzyme: sucrose-specific phosphotransferase system, EIIBC component (low affinity) <br/><br/> | + | * '''Description:''' [[trigger enzymes|trigger enzyme]]: sucrose-specific phosphotransferase system, EIIBC component (low affinity) <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 10: | Line 10: | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || trigger enzyme: sucrose-specific phosphotransferase system, | + | |style="background:#ABCDEF;" align="center"| '''Product''' || [[trigger enzymes|trigger enzyme]]: sucrose-specific phosphotransferase system, <br/>EIIBC component (low affinity) |
| − | EIIBC component (low affinity) | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || sucrose uptake and phosphorylation, control of [[SacY]] activity | |style="background:#ABCDEF;" align="center"|'''Function''' || sucrose uptake and phosphorylation, control of [[SacY]] activity | ||
Revision as of 19:47, 1 September 2009
- Description: trigger enzyme: sucrose-specific phosphotransferase system, EIIBC component (low affinity)
| Gene name | sacX |
| Synonyms | ipa-14r, sacS |
| Essential | no |
| Product | trigger enzyme: sucrose-specific phosphotransferase system, EIIBC component (low affinity) |
| Function | sucrose uptake and phosphorylation, control of SacY activity |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 48 kDa, 9.346 |
| Gene length, protein length | 1377 bp, 459 aa |
| Immediate neighbours | epr, sacY |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 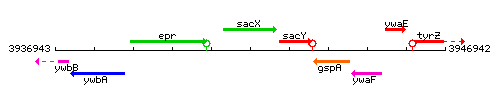
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU38410
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Protein EIIB N(pi)-phospho-L-histidine/cysteine + sugar = protein EIIB + sugar phosphate (according to Swiss-Prot)
- Protein family: PTS permease, sucrose permease (Scr) family PubMed
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P15400
- KEGG entry: [3]
- E.C. number: 2.7.1.69
Additional information
Expression and regulation
- Regulation: induction by sucrose (at high concentration)
- Regulatory mechanism: induction by SacY-dependent RNA switch (transcriptional antitermination)
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Jonathan Reizer, Steffi Bachem, Aiala Reizer, Maryvonne Arnaud, Milton H Saier, Jörg Stülke
Novel phosphotransferase system genes revealed by genome analysis - the complete complement of PTS proteins encoded within the genome of Bacillus subtilis.
Microbiology (Reading): 1999, 145 ( Pt 12);3419-3429
[PubMed:10627040]
[WorldCat.org]
[DOI]
(P p)
A M Crutz, M Steinmetz, S Aymerich, R Richter, D Le Coq
Induction of levansucrase in Bacillus subtilis: an antitermination mechanism negatively controlled by the phosphotransferase system.
J Bacteriol: 1990, 172(2);1043-50
[PubMed:2105292]
[WorldCat.org]
[DOI]
(P p)