Difference between revisions of "SucD"
| Line 72: | Line 72: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
** Inhibited by 2-oxoglutarate, ATP and NADH [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6T36-44C8RWC-SH&_user=5731894&_coverDate=01%2F01%2F1985&_rdoc=28&_fmt=high&_orig=browse&_srch=doc-info(%23toc%234938%231985%23998209998%23270526%23FLP%23display%23Volume)&_cdi=4938&_sort=d&_docanchor=&_ct=40&_acct=C000043105&_version=1&_urlVersion=0&_userid=5731894&md5=f14f4734123ab1177d7217cab6c7ce7d FEBS Letters] | ** Inhibited by 2-oxoglutarate, ATP and NADH [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6T36-44C8RWC-SH&_user=5731894&_coverDate=01%2F01%2F1985&_rdoc=28&_fmt=high&_orig=browse&_srch=doc-info(%23toc%234938%231985%23998209998%23270526%23FLP%23display%23Volume)&_cdi=4938&_sort=d&_docanchor=&_ct=40&_acct=C000043105&_version=1&_urlVersion=0&_userid=5731894&md5=f14f4734123ab1177d7217cab6c7ce7d FEBS Letters] | ||
| − | + | ** GTP is not accepted by the enzyme [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6T36-44C8RWC-SH&_user=5731894&_coverDate=01%2F01%2F1985&_rdoc=28&_fmt=high&_orig=browse&_srch=doc-info(%23toc%234938%231985%23998209998%23270526%23FLP%23display%23Volume)&_cdi=4938&_sort=d&_docanchor=&_ct=40&_acct=C000043105&_version=1&_urlVersion=0&_userid=5731894&md5=f14f4734123ab1177d7217cab6c7ce7d FEBS Letters] | |
* '''Interactions:''' | * '''Interactions:''' | ||
| Line 93: | Line 93: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' ''[[sucC]]''-''[[ | + | * '''Operon:''' ''[[sucC]]''-''[[sucD]]'' {{PubMed|11976317}} |
* '''Sigma factor:''' | * '''Sigma factor:''' | ||
| − | * '''Regulation:''' repressed by glucose (2.4-fold) ([[CcpA]]) | + | * '''Regulation:''' repressed by glucose (2.4-fold) ([[CcpA]]) {{PubMed|12850135}} |
| − | * '''Regulatory mechanism:''' [[CcpA]]: transcription repression | + | * '''Regulatory mechanism:''' [[CcpA]]: transcription repression {{PubMed|12850135}} |
* '''Additional information:''' | * '''Additional information:''' | ||
| Line 123: | Line 123: | ||
=References= | =References= | ||
| − | <pubmed>12850135 17218307 | + | <pubmed>12850135 17218307 11976317</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 16:24, 11 October 2009
- Description: succinyl-CoA synthetase (alpha subunit)
| Gene name | sucD |
| Synonyms | |
| Essential | no |
| Product | succinyl-CoA synthetase (alpha subunit) |
| Function | TCA cycle |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 31 kDa, 5.587 |
| Gene length, protein length | 900 bp, 300 aa |
| Immediate neighbours | sucC, dprA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 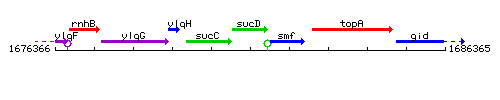
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU16100
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + succinate + CoA = ADP + phosphate + succinyl-CoA (according to Swiss-Prot)
- Protein family: succinate/malate CoA ligase alpha subunit family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: Reversible Michaelis-Menten FEBS Letters
- Domains:
- Modification: phosphorylation on (Ser-19 OR Thr-20) PubMed
- Cofactor(s):
- Effectors of protein activity:
- Inhibited by 2-oxoglutarate, ATP and NADH FEBS Letters
- GTP is not accepted by the enzyme FEBS Letters
- Interactions:
- Localization:
Database entries
- Structure: 1JKJ (E. coli)
- UniProt: P80865
- KEGG entry: [3]
- E.C. number: 6.2.1.5
Additional information
The enzyme is a dimer FEBS Letters
Expression and regulation
- Sigma factor:
- Additional information:
Biological materials
- Mutant: GP720 (spc), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References