Difference between revisions of "YrhJ"
| Line 78: | Line 78: | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * ''' | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/O08336 O08336] |
* '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU27160] | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU27160] | ||
Revision as of 13:50, 20 July 2009
- Description: similar to cytochrome P450 / NADPH-cytochrome P450 reductase
| Gene name | yrhJ |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | unknown |
| MW, pI | 118 kDa, 6.017 |
| Gene length, protein length | 3162 bp, 1054 aa |
| Immediate neighbours | yrhK, fatR |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 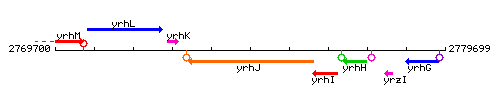
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU27160
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: NADPH + n oxidized hemoprotein = NADP+ + n reduced hemoprotein (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: O08336
- KEGG entry: [3]
- E.C. number: 1.6.2.4
Additional information
Expression and regulation
- Regulation: induced by unsaturated fatty acids (FatR)
- Regulatory mechanism: FatR: transcription repression
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Mattias C U Gustafsson, Olivier Roitel, Ker R Marshall, Michael A Noble, Stephen K Chapman, Antonio Pessegueiro, Armand J Fulco, Myles R Cheesman, Claes von Wachenfeldt, Andrew W Munro
Expression, purification, and characterization of Bacillus subtilis cytochromes P450 CYP102A2 and CYP102A3: flavocytochrome homologues of P450 BM3 from Bacillus megaterium.
Biochemistry: 2004, 43(18);5474-87
[PubMed:15122913]
[WorldCat.org]
[DOI]
(P p)
Oliver Lentz, Vlada Urlacher, Rolf D Schmid
Substrate specificity of native and mutated cytochrome P450 (CYP102A3) from Bacillus subtilis.
J Biotechnol: 2004, 108(1);41-9
[PubMed:14741768]
[WorldCat.org]
[DOI]
(P p)
M C Gustafsson, C N Palmer, C R Wolf, C von Wachenfeldt
Fatty-acid-displaced transcriptional repressor, a conserved regulator of cytochrome P450 102 transcription in Bacillus species.
Arch Microbiol: 2001, 176(6);459-64
[PubMed:11734890]
[WorldCat.org]
[DOI]
(P p)
T R Lee, H P Hsu, G C Shaw
Transcriptional regulation of the Bacillus subtilis bscR-CYP102A3 operon by the BscR repressor and differential induction of cytochrome CYP102A3 expression by oleic acid and palmitate.
J Biochem: 2001, 130(4);569-74
[PubMed:11574077]
[WorldCat.org]
[DOI]
(P p)