Difference between revisions of "PyrE"
| Line 80: | Line 80: | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * ''' | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/P25972 P25972] |
* '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU15560] | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU15560] | ||
Revision as of 12:43, 20 July 2009
- Description: orotate phosphoribosyltransferase
| Gene name | pyrE |
| Synonyms | pyrX |
| Essential | no |
| Product | orotate phosphoribosyltransferase |
| Function | pyrimidine biosynthesis |
| Metabolic function and regulation of this protein in SubtiPathways: Pyrimidines, Nucleotides (regulation) | |
| MW, pI | 23 kDa, 5.631 |
| Gene length, protein length | 648 bp, 216 aa |
| Immediate neighbours | pyrF, cysH |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 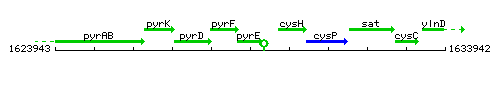
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU15560
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Orotidine 5'-phosphate + diphosphate = orotate + 5-phospho-alpha-D-ribose 1-diphosphate (according to Swiss-Prot)
- Protein family: PyrE subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: P25972
- KEGG entry: [3]
- E.C. number: 2.4.2.10
Additional information
Expression and regulation
- Regulatory mechanism:
- PyrR: RNA switch, transcription termination/ antitermination (in the presence of uridine nucleotides: termination, in their absence: antitermination) PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
R J Turner, Y Lu, R L Switzer
Regulation of the Bacillus subtilis pyrimidine biosynthetic (pyr) gene cluster by an autogenous transcriptional attenuation mechanism.
J Bacteriol: 1994, 176(12);3708-22
[PubMed:8206849]
[WorldCat.org]
[DOI]
(P p)
C L Quinn, B T Stephenson, R L Switzer
Functional organization and nucleotide sequence of the Bacillus subtilis pyrimidine biosynthetic operon.
J Biol Chem: 1991, 266(14);9113-27
[PubMed:1709162]
[WorldCat.org]
(P p)