Difference between revisions of "Eno"
| Line 75: | Line 75: | ||
* '''Interactions:''' [[Eno]]-[[PfkA]], [[Eno]]-[[Rny]] [http://www.ncbi.nlm.nih.gov/sites/entrez/19193632 PubMed] | * '''Interactions:''' [[Eno]]-[[PfkA]], [[Eno]]-[[Rny]] [http://www.ncbi.nlm.nih.gov/sites/entrez/19193632 PubMed] | ||
| − | * '''Localization:''' cytoplasm [http://www.ncbi.nlm.nih.gov/sites/entrez/16479537 PubMed] and membrane associated [http://www.ncbi.nlm.nih.gov/sites/entrez/18763711 PubMed] | + | * '''Localization:''' cytoplasm (according to Swiss-Prot), cytoplasm [http://www.ncbi.nlm.nih.gov/sites/entrez/16479537 PubMed] and membrane associated [http://www.ncbi.nlm.nih.gov/sites/entrez/18763711 PubMed] |
=== Database entries === | === Database entries === | ||
Revision as of 17:39, 30 April 2009
- Description: enolase, glycolytic/ gluconeogenic enzyme
| Gene name | eno |
| Synonyms | |
| Essential | yes |
| Product | enolase |
| Function | enzyme in glycolysis/ gluconeogenesis |
| MW, pI | 46,4 kDa, 4.49 |
| Gene length, protein length | 1290 bp, 430 amino acids |
| Immediate neighbours | pgm, yvgK |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 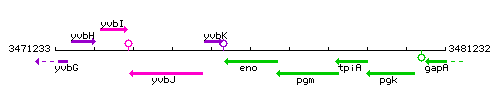
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates: 3475589 - 3476878
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2-phospho-D-glycerate = phosphoenolpyruvate + H(2)O
- Protein family: enolase family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- substrate binding domain (366–369)
- Cofactor(s): magnesium ion
- Effectors of protein activity:
Database entries
- Structure:
- Swiss prot entry: [3]
- KEGG entry: [4]
- E.C. number: 4.2.1.11
Additional information
Expression and regulation
- Sigma factor: SigA
- Regulation: expression activated by glucose (3.3 fold) PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector: pGP563 (N-terminal His-tag, in pWH844), pGP93 (N-terminal Strep-tag, for SPINE, in pGP380), available in Stülke lab
- lacZ fusion:
- GFP fusion: pHT315-yfp-eno, available in Mijakovic lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
- Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways. Metab Eng. 5: 133-149 PubMed
- Eymann et al. (2007) Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis. Proteomics 7: 3509-3526. PubMed
- Lévine et al. (2006) Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes. Proteomics 6: 2157-2173 PubMed
- Commichau, F. M., Rothe, F. M., Herzberg, C., Wagner, E., Hellwig, D., Lehnik-Habrink, M., Hammer, E., Völker, U. & Stülke, J. (2009) Novel activities of glycolytic enzymes in Bacillus subtilis: Interactions with essential proteins involved in mRNA processing. Mol. Cell. Proteomics in press PubMed
- Ludwig, H., Homuth, G., Schmalisch, M., Dyka, F. M., Hecker, M., and Stülke, J. (2001) Transcription of glycolytic genes and operons in Bacillus subtilis: evidence for the presence of multiple levels of control of the gapA operon. Mol Microbiol 41, 409-422.PubMed
- Jannière, L., Canceill, D., Suski, C., Kanga, S., Dalmais, B., Lestini, R., Monnier, A. F., Chapuis, J., Bolotin, A., Titok, M., Le Chatelier, E., and Ehrlich, S. D. (2007) Genetic evidence for a link between glycolysis and DNA replication. PLoS ONE 2, e447. PubMed
- Leyva-Vazquez, M. A., and Setlow, P. (1994) Cloning and nucleotide sequences of the genes encoding triose phosphate isomerase, phosphoglycerate mutase, and enolase from Bacillus subtilis. J Bacteriol 176: 3903-3910. PubMed
- Macek et al. (2007) The serine/ threonine/ tyrosine phosphoproteome of the model bacterium Bacillus subtilis. Mol. Cell. Proteomics 6: 697-707 PubMed