Difference between revisions of "DesK"
(→Original publications) |
|||
| (53 intermediate revisions by 6 users not shown) | |||
| Line 1: | Line 1: | ||
| − | * '''Description:''' two-component sensor kinase, regulation of cold shock expression of des <br/><br/> | + | * '''Description:''' [[two-component systems|two-component]] sensor kinase, regulation of cold shock expression of ''[[des]]'' <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 10: | Line 10: | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || two-component sensor kinase | + | |style="background:#ABCDEF;" align="center"| '''Product''' || [[two-component systems|two-component]] sensor kinase |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || regulation of cold shock expression of des | + | |style="background:#ABCDEF;" align="center"|'''Function''' || regulation of cold shock expression of ''[[des]]'' |
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU19190 desK] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=DesK DesK] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=desK desK]''' | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 42 kDa, 9.428 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 42 kDa, 9.428 | ||
| Line 20: | Line 26: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[des]]'', ''[[desR]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[des]]'', ''[[desR]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU19190 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU19190 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU19190 DNA_with_flanks] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:yocF_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:yocF_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=desK_2090574_2091686_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:desK_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU19190]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/> | ||
| + | |||
| + | = [[Categories]] containing this gene/protein = | ||
| + | {{SubtiWiki category|[[lipid metabolism/ other]]}}, | ||
| + | {{SubtiWiki category|[[protein modification]]}}, | ||
| + | {{SubtiWiki category|[[transcription factors and their control]]}}, | ||
| + | {{SubtiWiki category|[[cold stress proteins]]}}, | ||
| + | {{SubtiWiki category|[[membrane proteins]]}}, | ||
| + | {{SubtiWiki category|[[phosphoproteins]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| − | |||
=The gene= | =The gene= | ||
| Line 35: | Line 56: | ||
=== Basic information === | === Basic information === | ||
| − | * ''' | + | * '''Locus tag:''' BSU19190 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU19190&redirect=T BSU19190] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yocFG.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yocFG.html] | ||
| Line 46: | Line 68: | ||
=== Additional information=== | === Additional information=== | ||
| + | |||
| + | |||
| Line 52: | Line 76: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' autophosphorylation, phosphorylation of [[DesR]] |
| − | + | * '''Protein family:''' | |
| − | * '''Protein family:''' | ||
* '''Paralogous protein(s):''' | * '''Paralogous protein(s):''' | ||
| Line 63: | Line 86: | ||
* '''Domains:''' | * '''Domains:''' | ||
| + | ** 5 transmembrane helices | ||
| + | ** cytoplasmatic C-terminal trail | ||
| − | * '''Modification:''' | + | * '''Modification:''' autophosphorylation on a His residue |
* '''Cofactor(s):''' | * '''Cofactor(s):''' | ||
| − | * '''Effectors of protein activity:''' | + | * '''Effectors of protein activity:''' unsaturated fatty acids are negative effectors of the system |
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
| + | ** [[DesK]]-[[DesR]] | ||
| − | * '''Localization:''' | + | * '''[[Localization]]:''' membrane (transmembrane segments) |
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU19190&redirect=T BSU19190] | ||
| − | * '''Structure:''' | + | * '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=3EHF 3EHF] |
| − | * ''' | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/O34757 O34757] |
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU19190] |
* '''E.C. number:''' | * '''E.C. number:''' | ||
=== Additional information=== | === Additional information=== | ||
| + | *[[DesK]] has the ability to sense changes in membrane fluidity {{PubMed|17087771}} | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''[[desK]]-[[desR]]'' {{PubMed|11285232}} |
| − | * '''Sigma factor:''' | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=desK_2090574_2091686_1 desK] {{PubMed|22383849}} |
| + | |||
| + | * '''[[Sigma factor]]:''' | ||
* '''Regulation:''' | * '''Regulation:''' | ||
| + | ** induced by cold shock (12-fold) {{PubMed|12399512}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' |
=Biological materials = | =Biological materials = | ||
| Line 113: | Line 144: | ||
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| − | + | * [[Diego de Mendoza]], Universidad Nacional de Rosario, Argentine [http://www.ibr.gov.ar/eng/investigacion/demendoza.htm homepage] | |
| − | [[Mohamed Marahiel]], Marburg University, Germany [http://www.uni-marburg.de/fb15/fachgebiete/bio/marahiel?language_sync=1 homepage] | + | * [[Mohamed Marahiel]], Marburg University, Germany [http://www.uni-marburg.de/fb15/fachgebiete/bio/marahiel?language_sync=1 homepage] |
=Your additional remarks= | =Your additional remarks= | ||
=References= | =References= | ||
| + | ==Reviews== | ||
| + | <pubmed> 20117042, 17087771 24819366 </pubmed> | ||
| − | + | ==Original publications== | |
| + | <pubmed>10094672,14734164,12399512,20507988 , 19233289 11285232 19805278 15090506, 12207704 20705470 23356219 24574048 24522108 25406381 26172072 </pubmed> | ||
| + | [[Category:Protein-coding genes]] | ||
Latest revision as of 09:21, 15 July 2015
- Description: two-component sensor kinase, regulation of cold shock expression of des
| Gene name | desK |
| Synonyms | yocF |
| Essential | no |
| Product | two-component sensor kinase |
| Function | regulation of cold shock expression of des |
| Gene expression levels in SubtiExpress: desK | |
| Interactions involving this protein in SubtInteract: DesK | |
| Metabolic function and regulation of this protein in SubtiPathways: desK | |
| MW, pI | 42 kDa, 9.428 |
| Gene length, protein length | 1110 bp, 370 aa |
| Immediate neighbours | des, desR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 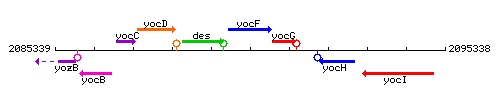
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed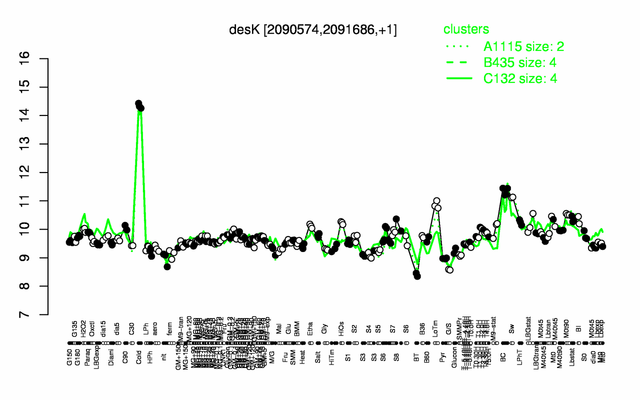
| |
Contents
Categories containing this gene/protein
lipid metabolism/ other, protein modification, transcription factors and their control, cold stress proteins, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU19190
Phenotypes of a mutant
Database entries
- BsubCyc: BSU19190
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: autophosphorylation, phosphorylation of DesR
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- 5 transmembrane helices
- cytoplasmatic C-terminal trail
- Modification: autophosphorylation on a His residue
- Cofactor(s):
- Effectors of protein activity: unsaturated fatty acids are negative effectors of the system
- Localization: membrane (transmembrane segments)
Database entries
- BsubCyc: BSU19190
- Structure: 3EHF
- UniProt: O34757
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- induced by cold shock (12-fold) PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Diego de Mendoza, Universidad Nacional de Rosario, Argentine homepage
- Mohamed Marahiel, Marburg University, Germany homepage
Your additional remarks
References
Reviews
Original publications