Difference between revisions of "DesK"
(→Reviews) |
|||
| Line 151: | Line 151: | ||
=References= | =References= | ||
==Reviews== | ==Reviews== | ||
| − | <pubmed> 20117042, 17087771</pubmed> | + | <pubmed> 20117042, 17087771 24819366 </pubmed> |
| + | |||
==Original publications== | ==Original publications== | ||
<pubmed>10094672,14734164,12399512,20507988 , 19233289 11285232 19805278 15090506, 12207704 20705470 23356219 24574048 24522108 </pubmed> | <pubmed>10094672,14734164,12399512,20507988 , 19233289 11285232 19805278 15090506, 12207704 20705470 23356219 24574048 24522108 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:14, 19 June 2014
- Description: two-component sensor kinase, regulation of cold shock expression of des
| Gene name | desK |
| Synonyms | yocF |
| Essential | no |
| Product | two-component sensor kinase |
| Function | regulation of cold shock expression of des |
| Gene expression levels in SubtiExpress: desK | |
| Interactions involving this protein in SubtInteract: DesK | |
| Metabolic function and regulation of this protein in SubtiPathways: desK | |
| MW, pI | 42 kDa, 9.428 |
| Gene length, protein length | 1110 bp, 370 aa |
| Immediate neighbours | des, desR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 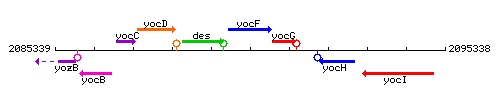
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed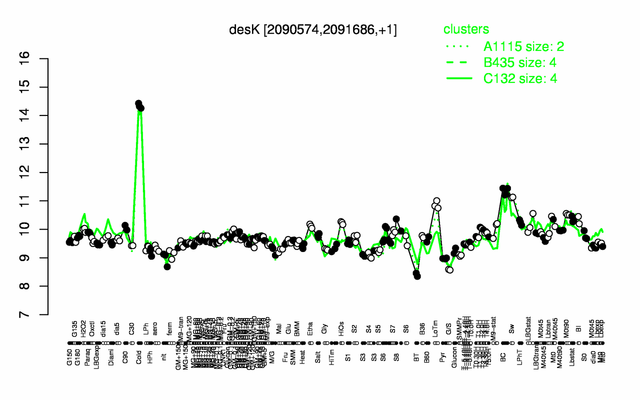
| |
Contents
Categories containing this gene/protein
lipid metabolism/ other, protein modification, transcription factors and their control, cold stress proteins, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU19190
Phenotypes of a mutant
Database entries
- BsubCyc: BSU19190
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: autophosphorylation, phosphorylation of DesR
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- 5 transmembrane helices
- cytoplasmatic C-terminal trail
- Modification: autophosphorylation on a His residue
- Cofactor(s):
- Effectors of protein activity: unsaturated fatty acids are negative effectors of the system
- Localization: membrane (transmembrane segments)
Database entries
- BsubCyc: BSU19190
- Structure: 3EHF
- UniProt: O34757
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- induced by cold shock (12-fold) PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Diego de Mendoza, Universidad Nacional de Rosario, Argentine homepage
- Mohamed Marahiel, Marburg University, Germany homepage
Your additional remarks
References
Reviews
Original publications