Difference between revisions of "HisC"
| (3 intermediate revisions by 2 users not shown) | |||
| Line 16: | Line 16: | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU22620 hisC] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU22620 hisC] | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/ | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=hisC hisC]''' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 39 kDa, 5.005 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 39 kDa, 5.005 | ||
| Line 55: | Line 55: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU22620&redirect=T BSU22620] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/trpEDCFBA-hisC-tyrA-aroE.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/trpEDCFBA-hisC-tyrA-aroE.html] | ||
| Line 89: | Line 90: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU22620&redirect=T BSU22620] | ||
* '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId=1UU0 1UU0] (the enzyme from ''E. coli'') {{PubMed|11518529}} | * '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId=1UU0 1UU0] (the enzyme from ''E. coli'') {{PubMed|11518529}} | ||
| Line 119: | Line 121: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
** the mRNA is substantially stabilized upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | ** the mRNA is substantially stabilized upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 2715 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 2096 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 4023 {{PubMed|21395229}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 2507 {{PubMed|21395229}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 6242 {{PubMed|21395229}} | ||
=Biological materials = | =Biological materials = | ||
| − | |||
* '''Mutant:''' | * '''Mutant:''' | ||
Latest revision as of 14:08, 17 April 2014
- Description: histidinol-phosphate aminotransferase / tyrosine and phenylalanine aminotransferase
| Gene name | hisC |
| Synonyms | aroJ |
| Essential | no |
| Product | histidinol-phosphate aminotransferase / tyrosine and phenylalanine aminotransferase |
| Function | biosynthesis of aromatic amino acids |
| Gene expression levels in SubtiExpress: hisC | |
| Metabolic function and regulation of this protein in SubtiPathways: hisC | |
| MW, pI | 39 kDa, 5.005 |
| Gene length, protein length | 1080 bp, 360 aa |
| Immediate neighbours | tyrA, trpA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 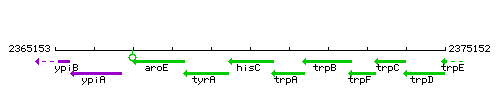
| |
Expression at a glance PubMed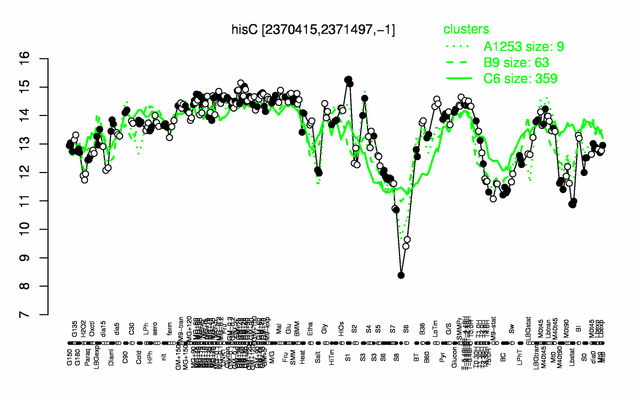
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22620
Phenotypes of a mutant
Database entries
- BsubCyc: BSU22620
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-histidinol phosphate + 2-oxoglutarate = 3-(imidazol-4-yl)-2-oxopropyl phosphate + L-glutamate (according to Swiss-Prot)
- Protein family: bacterial solute-binding protein 3 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU22620
- Swiss prot entry: P17731
- KEGG entry: [3]
- E.C. number: 2.6.1.9
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
- the mRNA is substantially stabilized upon depletion of RNase Y PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 2715 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 2096 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 4023 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 2507 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 6242 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References